AWS for Industries
Accelerate development of bioinformatics workflows on AWS HealthOmics using call caching
Bioinformatics workflows are computationally expensive, often running for hours to days. When these workflows fail mid-execution or require iterative refinement, rerunning from scratch wastes significant time and compute resources. AWS HealthOmics call caching solves this challenge by intelligently saving and reusing completed task outputs, enabling workflows to resume from failure points rather than restarting entirely. Call caching delivers four benefits: cost reduction by preventing unnecessary recompute, accelerated development cycles through reuse of computed results, production resilience via automatic failure recovery, and enhanced debugging through access to cached results.
In this blog, you will get an insight into how AWS HealthOmics call caching is setup and used to gain these benefits.
Overview
Genomics and multimodal analysis increasingly play an important role in applications like drug discovery, precision medicine, gene therapy, and agriculture. Due to the complex nature of data in these applications and sophisticated tools required to process them, bioinformatics workflows play a critical role in analyzing and interpreting them. These workflows can include a handful to hundreds of bioinformatics tools executed as part of a pipeline with complex dependencies, where some steps can be executed in parallel, while other steps need to be executed serially. These workflows often involve computationally intensive tasks that can run for extended periods from hours to multiple days to complete.
Workflows can fail due to reasons such as incorrect inputs, outdated tools and dependencies, infrastructure failures, or incremental development cycles. In these scenarios, re-running the entire workflow from scratch wastes valuable time and compute resources and extends workflow development cycles and processing turnaround times. AWS HealthOmics call caching solves this challenge by saving and reusing completed task outputs, reducing both runtime and costs.
Call caching features
Call caching is available for all three workflow languages currently supported by HealthOmics, i.e. Nextflow, Workflow Definition Language (WDL), and Common Workflow Language (CWL). HealthOmics uses Amazon Simple Storage Service (S3) as the backend to store cached data for workflow runs due to its durability, availability, cost-effectiveness, and virtually unlimited scalability. There are two caching behaviors supported:
- Cache on failure – task outputs from all completed tasks of a workflow run are cached only if the workflow fails. This is recommended for production runs where an edge case is encountered, and user intervention is needed to re-run from the last successful task.
- Cache always – task outputs from all tasks of a workflow run, irrespective of failure or success, are cached. This is recommended when developers want to rapidly iterate over a workflow and would like to fine-tune workflow code, docker images, and/or parameters of one or more tasks to achieve the desired results.
Note that intermediate files within tasks can be cached if they are declared as task outputs.
Caching requirements for tasks
HealthOmics caches task outputs for tasks that meet the following requirements:
- The task must define a container
- The task must produce one or more outputs
Additionally, if you suspect that a task produces outputs that are non-deterministic (such as random number generator or system time, or race-conditions that may cause output variance), consider not caching these specific tasks.
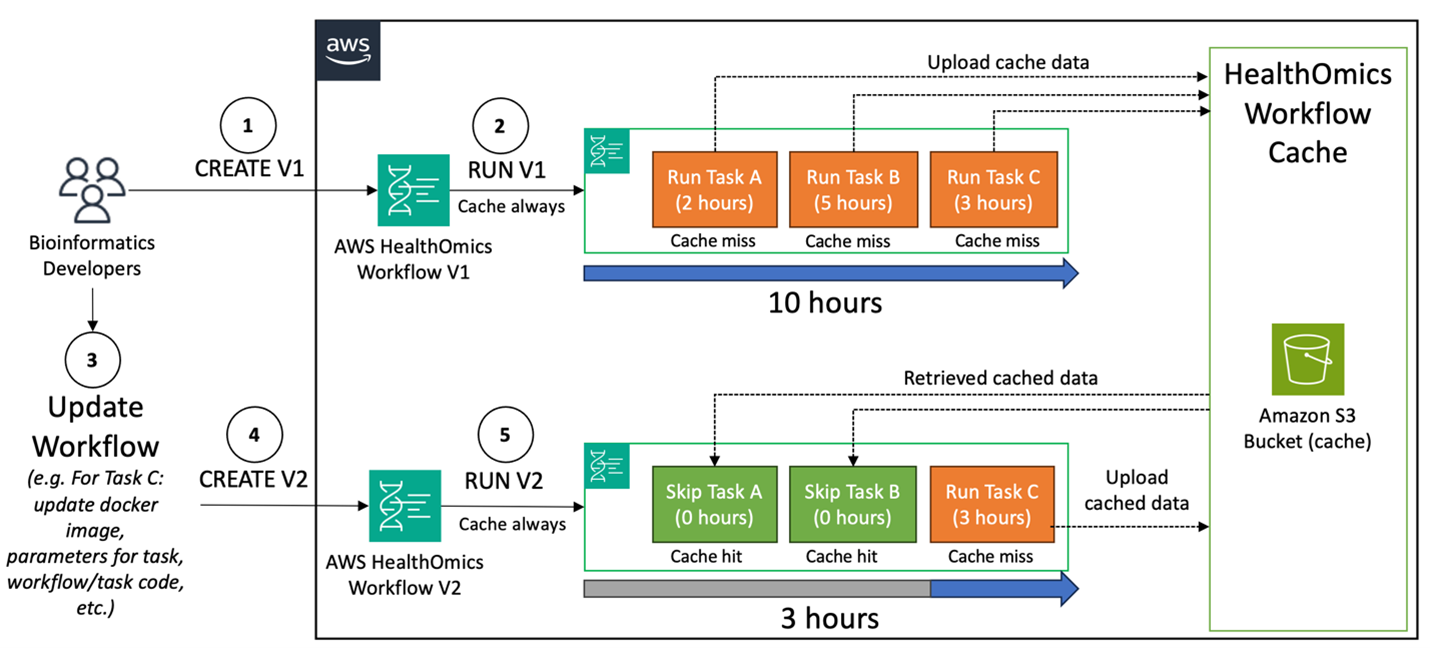
Figure 1: In this diagram, we show how cache always behavior during active workflow development can reduce the time between iterations and improve turnaround time to a finalized workflow.
Although HealthOmics provides a consistent caching experience across all three supported workflow languages and their engines, there are some engine specific features that are available that provide ability to disable task level call caching.
- For Nextflow, you can turn off caching for individual tasks by using the cache false directive. For information on this directive, see the Processes in the Nextflow specifications.
- For CWL, you can control caching for individual tasks by using the WorkReuse feature. For information on this directive, see the WorkReuse directive in CWL documentation.
- For WDL, you can disable caching for individual tasks by using the volatile attribute. For information on this directive, see the documentation on Cromwell website describing the Volatile optimization.
For additional engine-specific considerations you can visit the HealthOmics documentation.
Multiple caches can be created and used as per organizational needs. Note that users are responsible for the cost, data management, and lifecycle operations of the cached data in S3. Additional guidance on how to manage your run cache is available in the HealthOmics documentation.
Additionally, HealthOmics provides cache hit/miss status in the run details view as well as integrated CloudWatch logs which helps users understand how their changes to the workflow or inputs have impacted the underlying tasks. Users can access cache data in S3 to inspect intermediate files, engine logs, and artifacts. to debug errors and dive deeper. Additional details and considerations are available in the HealthOmics user guide.
How to use call caching within AWS HealthOmics
The following steps describe how you can set up and use the call caching feature within AWS HealthOmics workflows:
- Users create a cache and provide an existing Amazon S3 bucket as the backend storage for the cache. Users specify the default cache behavior – “Cache Always” or “Cache on failure”. Note that this behavior can be changed at the run level when a workflow is launched.
- Once the caches are created, users can follow their usual process to run a workflow with HealthOmics. Users can specify if the run should use a cache and can choose the cache behavior for this run.
- If the workflow run fails, the user can view the failure reason in HealthOmics, or inspect the logs available within HealthOmics via Amazon CloudWatch to find the reason for the failure. More details on ways to troubleshoot workflow runs are available in the HealthOmics documentation. Additionally, the user can access the cached task outputs in S3 to help debug or dive deeper into task executions details.
- The user updates the workflow component that caused the failure, such as, a particular task’s docker image, a specific task’s resources, the workflow code for the error causing task, run parameters if the inputs were incorrect, etc. Once the workflow is updated, the user can create a new HealthOmics workflow version.
- The user can now run the new workflow version with the cache used by the previous workflow’s run.
- On run completion, the tasks that previously completed in the original workflow and were not updated in this version will not be executed. Instead, their results will be retrieved from the cache, and the workflow will only run the tasks impacted by the change. The image below shows the “Cache hit” column links to the S3 location for the cached data for the task that has a cache hit.
Best Practices
- Cache structure and organization: Design caches around logical project boundaries (e.g. one cache per project), simplifies management and maximizing its reuse across workflows and workflow versions.
- Design Deterministic Tasks: Ensure your workflow tasks are idempotent – producing identical outputs when given identical inputs. Avoid system time dependencies, random number generators without fixed seeds, and concurrency-related race conditions, ensuring call caching is used appropriately
- Optimize Cache Storage Management: Establish regular cleanup schedule (e.g. end of major development project), monitor cache size regularly and clean up outdated cache entries, automate cache management through s3 lifecycle policies for retention, archiving or deleting based on your development cycle.
- Cache intermediate files strategically: Declare important intermediate files as task outputs to enable advanced debugging
Key Benefits
- Cost Optimization: Genomic workflows can consume significant compute resources. Call caching eliminates redundant computation by reusing results from previous runs, directly reducing your compute costs.
- Accelerated Development Cycles: During workflow development, researchers frequently need to modify and test specific pipeline components. Instead of rerunning entire workflows, call caching allows you to iterate on individual tasks while reusing outputs from unchanged components, dramatically speeding up the development process.
- Resilient Production Workflows: Production genomic workflows processing critical samples can fail due to various factors – infrastructure issues, data problems, or resource constraints. Call caching enables automatic resumption from the point of failure, ensuring that successfully completed tasks don’t need to be recomputed.
- Debugging and Troubleshooting: By caching intermediate files as task outputs, researchers gain access to execution artifacts for debugging. This visibility into intermediate processing steps is useful for troubleshooting complex genomic analysis pipelines.
Conclusion
By eliminating the need to completely rerun previously executed workflow tasks, call caching in AWS HealthOmics helps customers accelerate scientific breakthroughs by reducing both computational costs and time-to-insight for omics analyses. Bioinformaticians can rapidly iterate to develop accurate and performant workflows whilst being cost efficient. Consider leveraging call caching within AWS HealthOmics to accelerate your bioinformatics analysis needs. Contact your AWS account representatives or visit the AWS HealthOmics webpage to learn more about AWS HealthOmics and related resources.
Additional Resources
- AWS HealthOmics Official Documentation – Comprehensive guide covering HealthOmics concepts, features, storage, analytics, and workflows with getting started tutorials
- AWS HealthOmics Tutorials and Examples – Hands-on tutorials and code samples for working with HealthOmics workflows, storage, and analytics
- Run Optimization and Cost Analysis Blog – In-depth guide on using HealthOmics Run Analyzer for optimizing workflow performance and reducing costs
- Multi-Modal Data Analysis Guidance – End-to-end framework showing how to integrate HealthOmics with other AWS health and ML services
- AWS HealthOmics Call Caching Documentation – Complete guide to call caching concepts, implementation, and best practices for optimizing workflow reruns
- AWS HealthOmics MCP Server – Documentation for the Model Context Protocol server specifically designed for AWS HealthOmics integration.