AWS for Industries
Introducing Amazon Bio Discovery
A new agentic application that makes lab-in-the-loop drug discovery accessible to every researcher
Lab-in-the-loop drug discovery has transformed research for some organizations. AI-powered predictions improve continuously through wet-lab feedback, accelerating the path from hypothesis to validated candidates. But for most research teams, the reality looks different.
The field is moving fast. New biological AI models emerge constantly, each with different strengths, data requirements, and integration needs. Computational biologists are expected to evaluate and operationalize these models while supporting a growing number of discovery programs, often without the infrastructure or resources to match the demand. Meanwhile, bench scientists bring deep biological expertise to their targets and experiments but lack direct access to the computational tools that could accelerate their work. The result is a collaboration bottleneck: not because the science isn’t available, but because the tooling doesn’t support how these teams need to work together.
Even when systems are running, computational predictions and wet-lab workflows stay disconnected. Manual handoffs introduce delays, make it harder to reproduce experiments, and slow the feedback loop that makes lab-in-the-loop valuable in the first place. Scaling across multiple discovery programs and research teams remains a persistent challenge.
Amazon Bio Discovery changes this by bringing computational design and wet-lab validation together in one application. It makes lab-in-the-loop accessible and scalable across your entire research organization.
The application provides access to 40+ AI biology models with AI-guided selection. Users can also upload custom models as well as models licensed from third parties. Agentic assistants help you select the right models for your research goals, optimize configurations, and evaluate candidates for experimentation. Amazon Bio Discovery’s contract research organization (CRO) partners enable seamless wet-lab validation, with results flowing back to improve the next cycle.
For computational biologists, this means building, modifying, and enhancing computational workflows in a no-code environment without managing infrastructure or provisioning compute for training and inference. You can ensure your workflows have standardized data processing and rigorous analysis built in, then publish them for your team to use. For bench scientists, it means running multiple experiment versions in parallel and adjusting input parameters through agentic assistance, rather than waiting for someone to build a custom solution. Both roles work from the same system, the same data, and the same results.
This is where collaboration compounds. Computational biologists create reusable workflows that embed and scale their expertise in model selection, pipeline design, and analytical rigor, encoding decisions like which AI biology models to chain together, how to process and validate input data, and what quality thresholds to enforce. Bench scientists then pick up those workflows and apply their specialized knowledge of target biology and experimental context to configure experiments, evaluate computational results, and send validated candidates to the wet lab for testing. The results flow back to refine the models, and with each cycle, the workflows become more accurate, computational biologists can support even more programs, and bench scientists gain access to increasingly powerful tools. Projects that would have waited in a queue move forward immediately, and the value multiplies over time.
Figure 1: Begin creating a new recipe by choosing your workflow type in a no-code environment
See it in Action
Let’s walk through a representative antibody design workflow to show you what this looks like in practice. This mirrors the approach used in our collaboration with Memorial Sloan Kettering Cancer Center (MSK) where Amazon Bio Discovery was used to design nearly 300,000 novel antibody candidates, filter down to the top 100,000, and send them to the wet lab for testing in weeks versus up to a year using traditional design methods. Read the white paper to learn more.
Step 1. Evaluate models and build your in silico workflow
Start by exploring the catalog of 40+ AI biology models, including any models you have uploaded yourself. Each is specialized for different aspects of antibody design, from binding affinity prediction to developability assessment. The module evaluation page lets you filter by the property you’re interested in. You can run head-to-head comparisons using our antibody developability benchmark. Read the blog to learn more about the benchmark dataset.
Not sure which models to use? The AI-assisted workflow walks you through what you want to achieve. You’ll see a list of recommendations across all hosted models, complete with scientific rationale. Review the suggestions, confirm your selections, and the system generates a recipe. This is a computational workflow pipeline you can modify in the sandbox and then publish for use in your experiments or share with colleagues.
This enables computational biologists to focus on advancing science rather than infrastructure. Instead of building individual pipelines or writing custom code for each project, you’re creating reusable workflows that anyone on your team can run. The computational expertise gets embedded in the recipe. The biological expertise drives how it’s used. Projects that would have waited in a queue can move forward immediately.
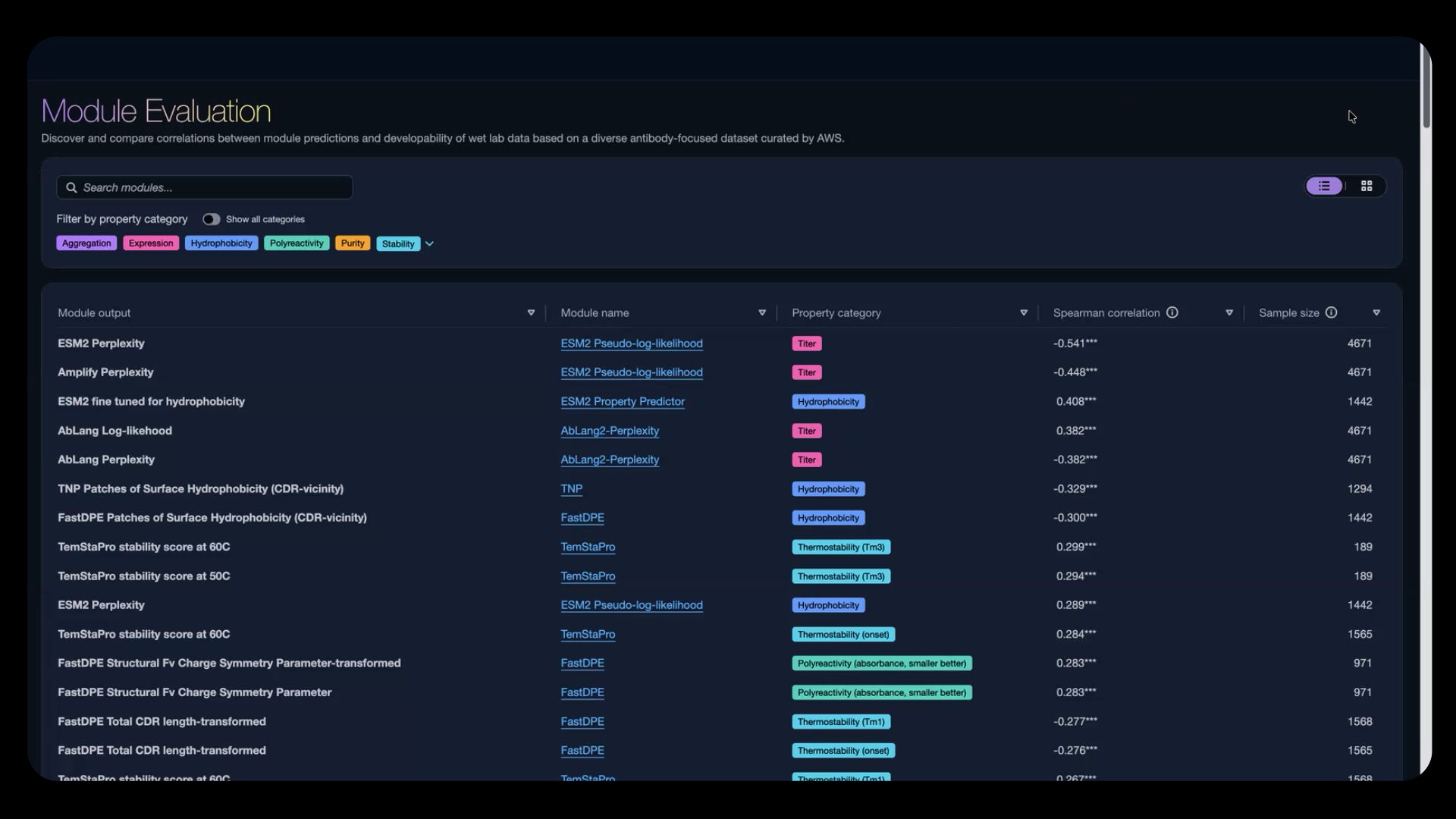
Figure 2: Benchmark AI biology model developability performance against real antibody dataset
Step 2: Design your in silico experiment with configuration agents
Once you’re ready to run an experiment, the AI agent guides you through the key decisions. For antibody design, that means identifying hotspot residues (the binding-critical amino acids) and selecting the right framework, the structural scaffold that affects how well your antibody binds to the target antigen.
The agent searches through multiple data sources and considers factors like surface accessibility and hydrophobicity. It provides recommendations with scientific rationale and references. You’re not just getting predictions. You’re understanding why the system is suggesting what it’s suggesting. If you want to adjust parameters or explore alternative approaches, you can do that directly. No coding required.
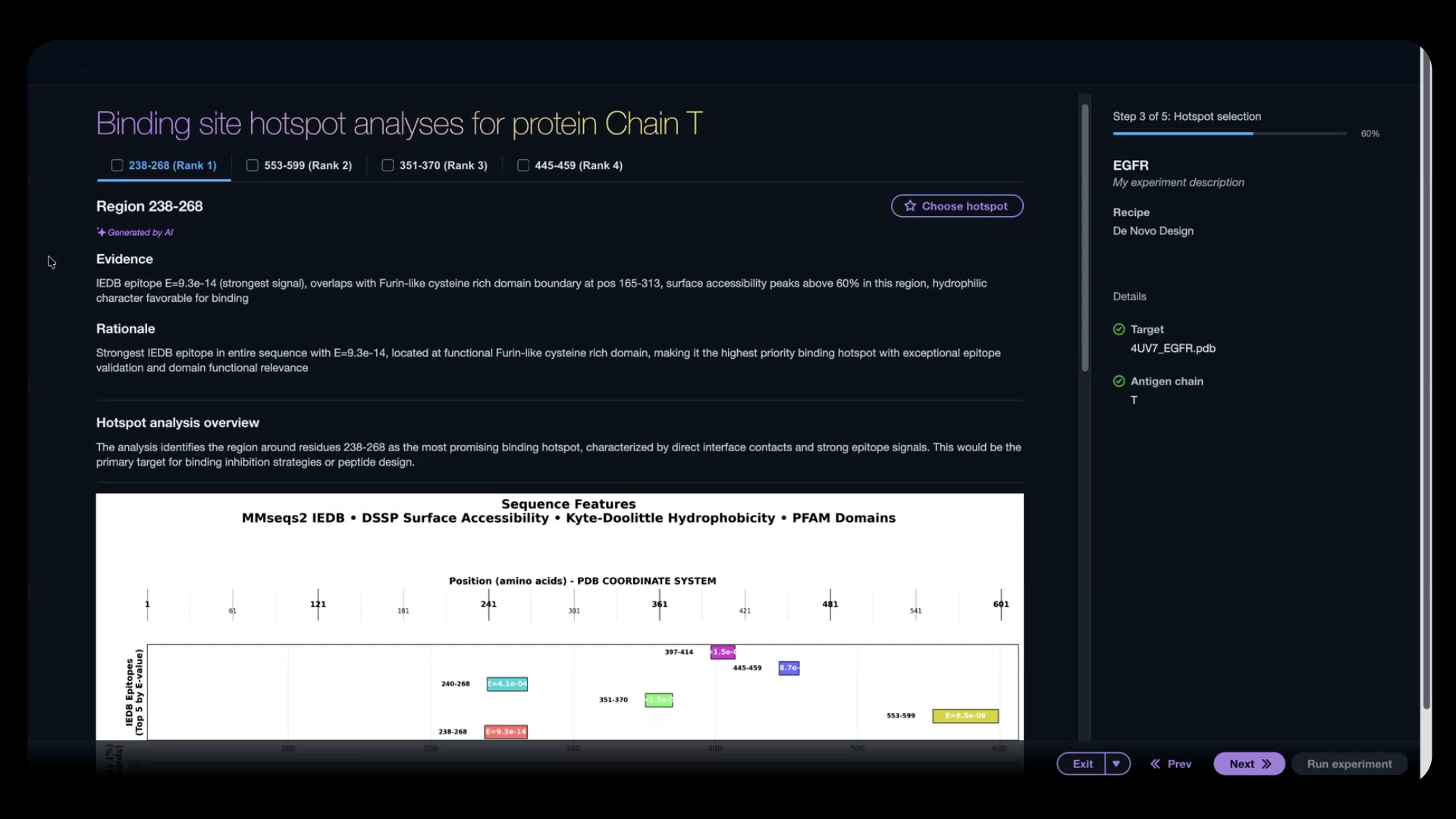
Figure 3: The configuration agent identifies hotspot residues on your target antigen
Step 3: Analyze in silico results with candidate selection agents
When the in silico experiment completes, you’ll see an AI-generated summary of your results and a pre-filtered list of candidates selected for you to consider. These candidates have already been run through multi-property optimization and liability assessment. This ensures no chemical modifications that would impact antibody stability, efficacy, or developability.
Each recommended candidate comes with rationale explaining why it was selected. You also have advanced analytical tools at your fingertips. Molecular dynamics analysis, functional and sequential diversity reviews let you dig into the data before deciding which candidates to move forward. The system gives you the insights you need to make more informed decisions faster.
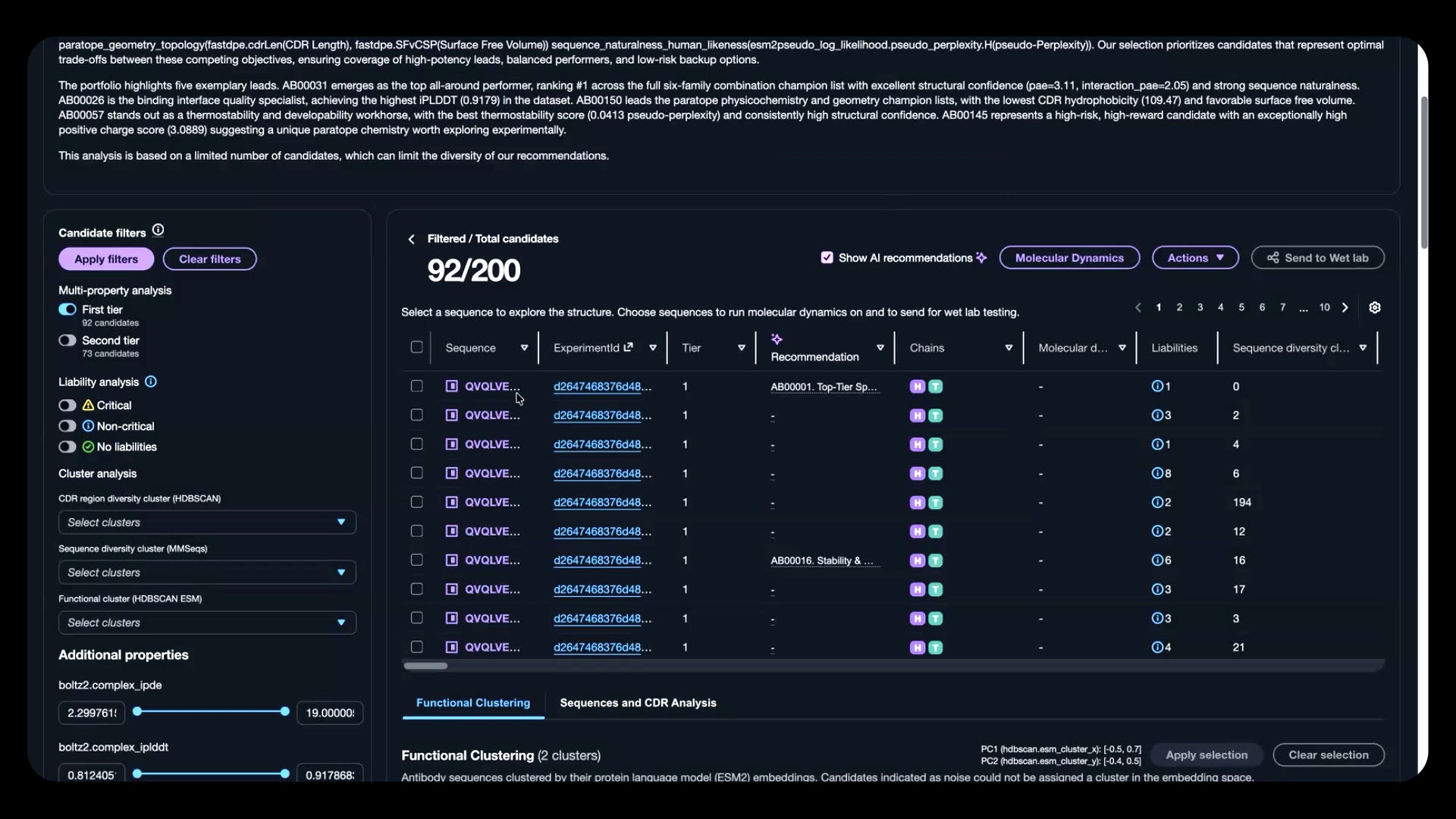
Figure 4: Review AI-recommended candidates filtered by multi-property optimization and liability assessment
Step 4: Send top candidates to wet lab for validation
Once you’ve identified a cohort of candidates, you can send them directly to one of our integrated lab partners: Ginkgo Bioworks, Twist Bioscience, or A-Alpha Bio. Select the assays you want to run, and you’ll get cost and turnaround time estimates in near real-time. No more manual handoffs. No more custom integrations with each CRO’s unique offerings.
When lab testing completes, the data flows back into Amazon Bio Discovery automatically. You can download results for further analysis or compare in silico predictions against wet-lab outcomes to understand prediction accuracy. The data triage and wrangling that used to require manual solutions is now handled through an experimental data registry—one place to view experimental inputs and results, now and in the future.
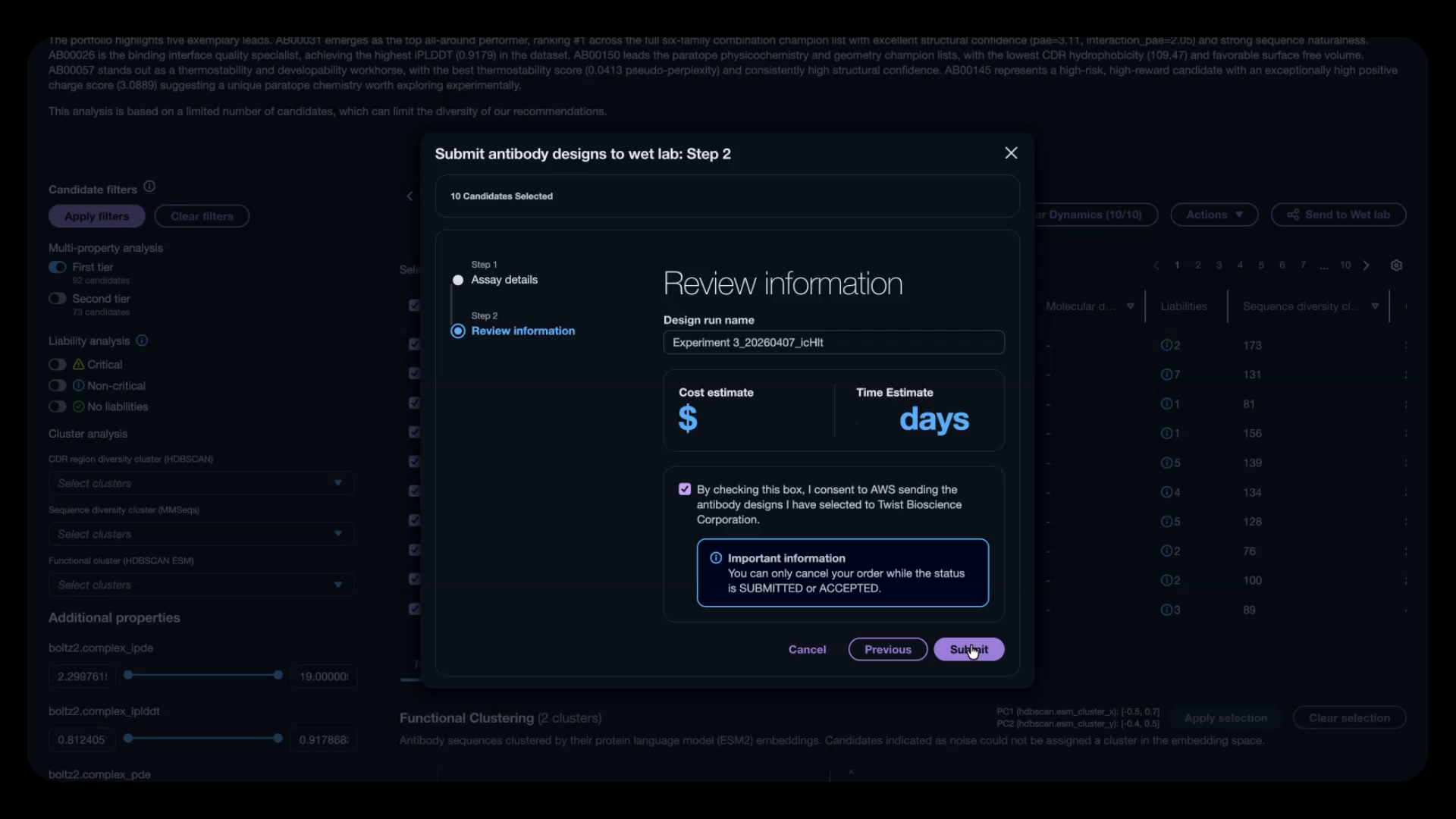
Figure 5: Send validated candidates directly to an integrated CRO partner with assay details, cost estimates, and turnaround times
Step 5: Close the loop with active learning
Here’s where lab-in-the-loop becomes real. The wet-lab results feed back into the system, allowing you to fine-tune models with your newly generated data. After a few cycles, you’re identifying drug-like candidates with increasing confidence. These are candidates you’re ready to take to the next stage, whether that’s animal testing or pre-clinical development.
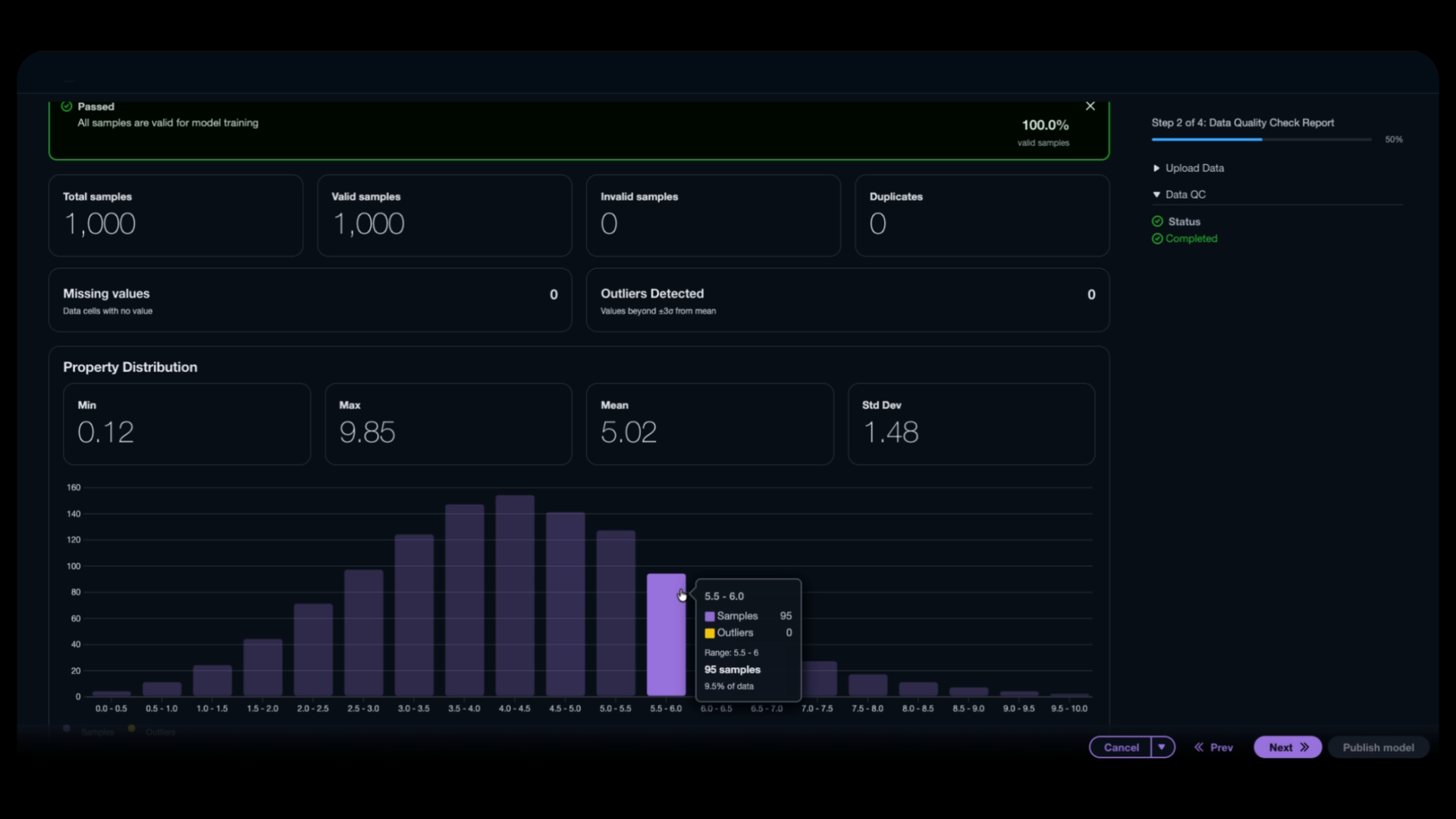
Figure 6: Verify incoming wet-lab data quality before model fine-tuning
Built for the way you work
Amazon Bio Discovery is built on the same AWS infrastructure trusted by 19 of the top 20 pharmaceutical companies. You get enterprise-grade security with data isolation to ensure your proprietary experimental data and custom-trained models remain protected within your application environment.
Get Started
Lab-in-the-loop drug discovery shouldn’t take time away from science. It should be accessible to everyone doing the science, whether you’re designing the computational workflows or running the experiments that test your hypotheses.
Amazon Bio Discovery is available today. Visit the Amazon Bio Discovery website to learn more, sign up for a free trial, and build skills through a free digital course from AWS Training and Certification.