AWS for Industries
How Carlsberg’s Traitomic business leveraged AWS HealthOmics to power genetic trait development
Introduction
Traitomic, a spinoff from the Carlsberg Research Laboratory, develops better genetics for the food industry. Traitomic uses technology developed at Carlsberg to help food companies create healthier, more sustainable products by identifying improved traits in crops and microbes, from higher-protein beans to more energy efficient microbial strains.
This blog explains how Traitomic partnered with AWS to power genetic trait development by overcoming common challenges with sequencing workflows.
Sequencing Challenges in Life Sciences
Life science organizations rely on genomic sequencing to solve critical problems across research, agriculture, and healthcare. However, the path from raw sequencing data to actionable insights presents significant technical hurdles.
Common Sequencing Problems
Organizations use sequencing to address diverse challenges:
- Trait discovery and breeding: Identifying genetic markers for desired characteristics in crops or livestock
- Variant analysis: Detecting mutations linked to disease or drug resistance
- Microbial characterization: Understanding strain properties for fermentation, probiotics, or food safety
- Population genomics: Analyzing genetic diversity across species or patient cohorts
- Quality control: Verifying genetic identity and purity in production strains or seed lines
Technical Challenges
The complexity of genomic analysis creates three major bottlenecks:
Data management: A single sequencing run can generate terabytes of data. Organizations struggle with storage costs, data transfer speeds, and maintaining data integrity across analysis pipelines. Many lack the infrastructure to efficiently move data from sequencers to compute environments, creating delays that slow research timelines.
Limited infrastructure: High-performance computing requirements for sequence alignment, variant calling, and comparative genomics often exceed on-premises capabilities. Organizations face difficult choices: invest heavily in local infrastructure that may sit idle between projects or accept the limitations of underpowered systems that extend analysis times from hours to days.
Required technical skills: Genomic analysis demands expertise spanning bioinformatics, computational biology, and cloud infrastructure. Teams need to understand both biological questions and technical tools like workflow languages, containerization, and pipeline orchestration. This skill gap is particularly acute at smaller organizations or those new to large-scale sequencing.
These challenges compound as sequencing volumes grow. What works for occasional projects becomes unsustainable at scale, forcing organizations to rethink their approach to genomic data analysis.
As an organization working with sequencing data, Traitomic has experienced similar challenges:
- Compute constrained in scale by on-premises availability
- Difficulty managing large data volumes and workflows written in different definition languages
- Overspending on static infrastructure
- An IT architecture that required their scientists to be both technical and domain experts
The Solution
To address these challenges, Traitomic adopted an AWS-hosted solution built on GitHub Actions, AWS HealthOmics, Amazon Simple Storage Service (S3), and WinSCP/Cyberduck for production sequencing workflows. The team is also evaluating Kiro-CLI and the HealthOmics MCP server for future production use. Together, these technologies deliver several key improvements:
AWS HealthOmics provides cost and scale improvement with pay-as-you-go managed bioinformatics infrastructure that provisions compute and storage when a workflow starts and de-provisions the infrastructure at workflow completion.
GitHub Actions decouples backend processing of sequencing data from a GitHub interface familiar to scientists. Triggering on-demand HealthOmics workflow runs from form submissions through GitHub allows scientists to focus on their research and engineers to focus on the supporting technology.
Amazon S3 stores terabytes of data cost efficiently using tiered storage to provide the scale needed for sequencing workloads. Third party transfer clients like WinSCP and Cyberduck provide an agentless drag-and-drop UI that allow scientists to easily move data to the cloud using Amazon S3 APIs, with limited overhead on regulated and latency-sensitive lab machines.
Kiro-CLI enables automated conversion of workflow definition files from Snakemake to NextFlow, WDL, or CWL using conversational AI. When integrated with the HealthOmics MCP server, Kiro-CLI provides additional capabilities such as workflow generation, validation, packaging, and troubleshooting.
The Architecture
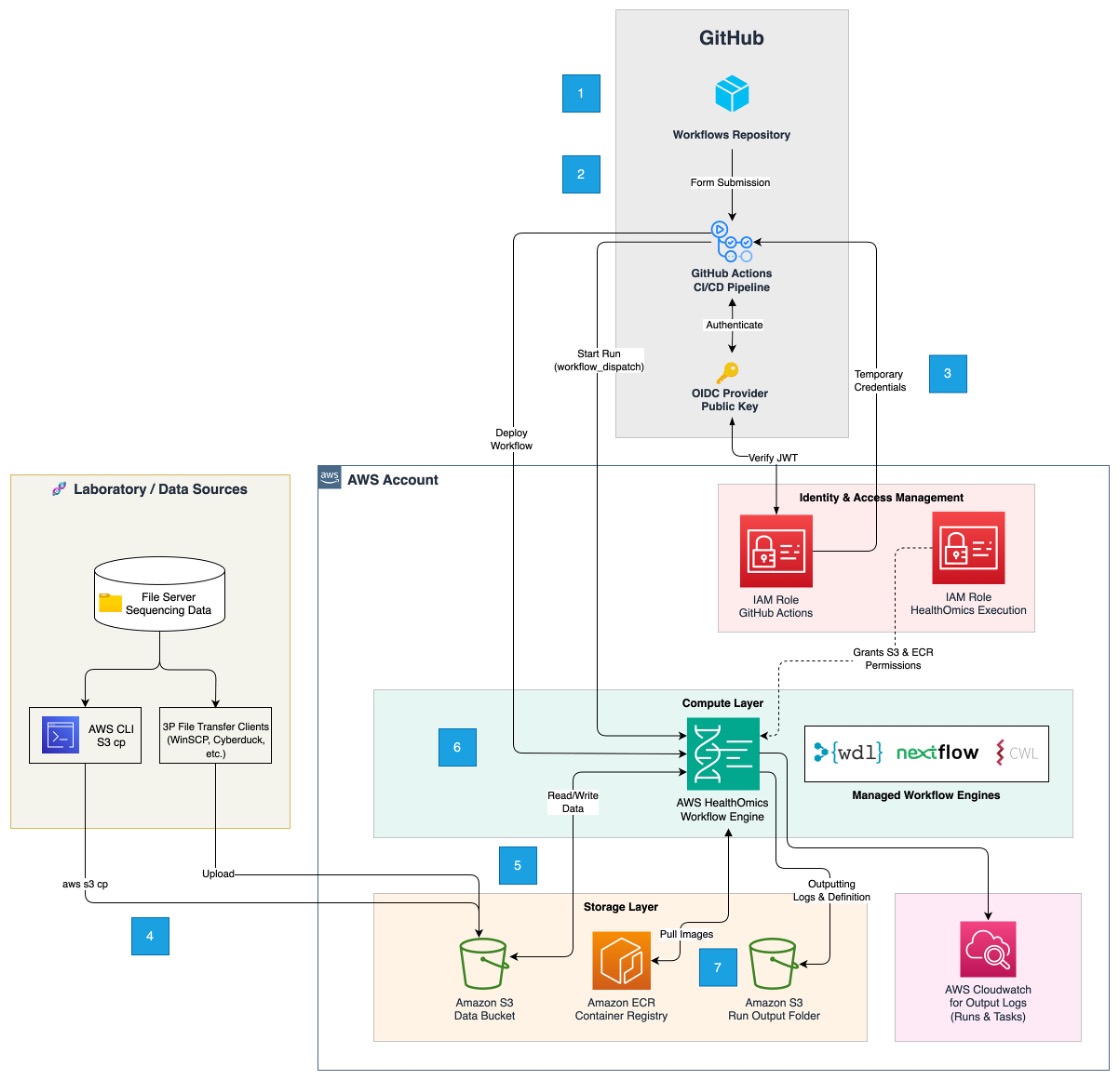
Figure 1: Traitomic’s sequencing architecture. Steps are described below.
- Bioinformaticians build HealthOmics workflows using a workflow definition language like NextFlow, WDL, or CWL. These workflows are pushed to GitHub for collaborative use. A GitHub Actions pipeline uses these workflow definitions to create and store the workflows in HealthOmics.
- Scientists log into GitHub. To use the workflows previously developed, they trigger a separate GitHub Actions workflow by submitting inputs like run_name, output_bucket, output_folder, storage_type, and workflow_name into a form through the GitHub UI.
- GitHub Actions interacts with AWS Identity Access Management to exchange an OpenID Connect token with the short-term credentials needed to invoke a HealthOmics run.
- WinSCP and Cyberduck allow on-demand migration of sequencing data from the lab to S3 so it’s available for use with HealthOmics.
- GitHub Actions invokes a HealthOmics run using the parameters specified in the GitHub UI. Required input and reference data is pulled from S3 privately by HealthOmics when the run is called. This data is stored in a temporary filesystem that’s only accessible to HealthOmics.
- HealthOmics dynamically provisions infrastructure to process each task in a workflow. When the task is completed, the infrastructure is deprovisioned for cost-optimization.
- Workflow outputs are stored in S3 where they are staged for downstream analysis.
Business Outcomes
Since deploying this solution, Traitomic engineers have received feedback that their scientist teams prefer working with HealthOmics compared to the on-premises HPC cluster. This is because the interface for HealthOmics is intuitive and because HealthOmics-managed pipeline orchestration allows scientists to perform more research by minimizing time spent monitoring active runs. Further, the time reduction for initiating and completing runs has allowed Traitomic to provide faster turnaround times to their customers. Consequently, Traitomic is migrating all their workflows to HealthOmics to scale these benefits.
Conclusion
Traitomic was able to increase the scale and usability of their bioinformatics pipelines by choosing HealthOmics over on-prem HPC infrastructure. To learn more about overcoming sequencing challenges with HealthOmics, read more about other production use cases. To get started with HealthOmics today, visit the HealthOmics GitHub repository for self-service tutorials exploring leveraging HealthOmics for sequencing data storage, workflows, and analytics.