AWS for Industries
Submit up to 100,000 Bioinformatics Workflow Runs with a Single API Call in AWS HealthOmics
AWS HealthOmics is a fully managed bioinformatics service designed to accelerate scientific breakthroughs at scale. Researchers and bioinformatics teams use HealthOmics to run complex workflows for drug discovery, population genomics studies, and clinical research applications.
Bioinformatics and drug discovery studies often need to process thousands or tens of thousands of samples through the same workflow with nearly identical parameters. Previously, the HealthOmics StartRun API supported up to five transactions per second (TPS), and runs had to be submitted individually. Researchers with thousands of runs would need to build submission scripts with loops and sleep intervals, or implement complex retry logic with exponential backoff and jitter strategies, or else receive HTTP 429 “Too Many Requests” errors, indicating rate limit violations. You also had to track the IDs of all runs in the batch, and complex logic was required if you needed to cancel or delete a batch.
This orchestration diverted valuable engineering time to infrastructure management instead of scientific discovery. In this blog post you will learn how the new HealthOmics batch runs can solve this problem for you.
The Solution
We launched AWS HealthOmics batch runs—a capability that transforms how you submit large-scale workflow processing jobs. With the StartRunBatch API, you can submit up to 100,000 runs in a single API request.
Batch runs allow you to define a common base configuration once and apply it across all runs in the batch. Each run can have different inputs and run-specific parameters through per-run overrides. This approach reduces submission overhead and simplifies lifecycle management for large-scale workflow processing.
The batch runs feature provides several key capabilities:
Unified Configuration Management: Define shared parameters such as workflow ID, IAM role, output URI, and common workflow parameters once in the defaultRunSetting. Individual runs inherit this configuration while allowing overrides for sample-specific values like input files, output locations, priority levels, and tags.
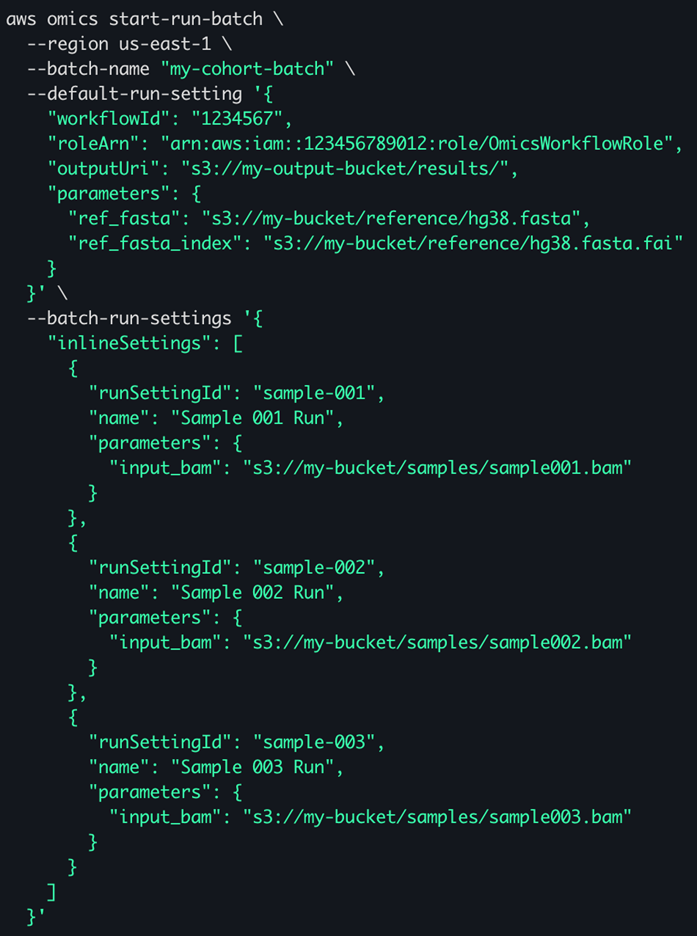
Flexible Submission Options: For batches up to 100 runs, you can provide configurations directly in the API request using inline settings (figure 1). For larger batches, store your run configurations in a JSON file in Amazon S3 and reference it in the API call. This approach supports the full 100,000 run limit (figure 2).
Figure 1 – an example start-batch-run command showing in-line –batch-run-settings for three samples. Settings used by all runs in the batch are specified with the –default-run-setting parameter.
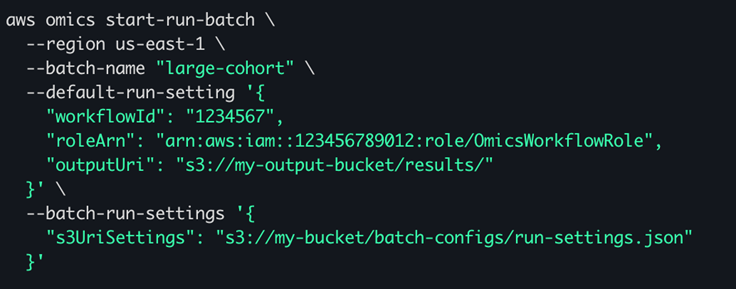
Figure 2 – an example start-batch-run command. The –batch-run-settings parameter references a JSON object in S3 which may contain up to 100,000 run settings.
Gradual Asynchronous Submission: HealthOmics validates your batch configuration synchronously and returns a batch ID. Individual runs are then submitted gradually and asynchronously at a controlled rate according to your account quotas. This eliminates the need for custom throttling logic in your code.
Comprehensive Monitoring: You can track overall batch status and individual run submission progress through the GetBatch and ListRunsInBatch APIs. Each run configuration includes a customer-provided runSettingId that maps to the HealthOmics-generated runId, allowing you to trace results back to your input samples.
Batch Lifecycle Management: You can: cancel all runs in a batch with a single CancelRunBatch call, delete all runs using DeleteRunBatch, and remove batch metadata with DeleteBatch. These operations simplify cleanup and resource management at scale.
The Benefit
Previously, submitting and managing large batches of runs required complex orchestration logic. Now you can achieve this with a single API call. Development teams can eliminate hundreds of lines of submission and retry code, reducing maintenance burden and potential error sources.
The impact extends beyond operational efficiency. By removing infrastructure complexity, researchers can focus on scientific questions rather than workflow orchestration. Faster submission times mean quicker turnaround for large cohort studies, accelerating the path from data to discovery.
Liz Baldo, Senior Staff Software Engineer at Manifold AI, shared her perspective: “This feature will significantly reduce our code complexity by having HealthOmics abstract away the complexity of batch workflow submissions. This will enable our users to seamlessly scale up their analyses on our platform.”
Organizations that process large sample cohorts—whether for population genomics, clinical trials, or drug discovery programs—gain immediate productivity improvements. The simplified API surface reduces onboarding time for new team members and decreases the likelihood of submission errors that could delay critical research timelines.
Conclusion
AWS HealthOmics batch runs eliminate the operational complexity of large-scale workflow submission. By consolidating up to 100,000 run submissions into a single API call with built-in retry logic and comprehensive monitoring, this feature allows your team to focus on scientific outcomes rather than infrastructure management. The combination of shared configuration management, flexible submission options, and simplified lifecycle operations provides a complete solution for production-scale genomics workflows.
Next Steps
You can create and manage HealthOmics batch runs today using the AWS command line interface (CLI), AWS software development kits (SDK), the HealthOmics Kiro power, and the HealthOmics MCP server.
Resources:
- Read the complete documentation: AWS HealthOmics Batch Runs User Guide
- Review the API reference: StartRunBatch API Documentation
- Explore HealthOmics: AWS HealthOmics Product Page
Start scaling your genomics workflows with batch runs and experience the difference that simplified submission makes for your research pipeline.