AWS for Industries
How to Connect AWS HealthOmics to Public and Private Network Sources at Runtime
Bioinformatic workflows often depend on connectivity to data sources like reference databases, variant annotation services, and public repositories such as NCBI and Ensembl. Previously, AWS HealthOmics workflows were executed in an isolated compute environment only able to access Amazon Simple Storage Service (S3) and HealthOmics storage. Now with Virtual Private Cloud (VPC) connected workflows, AWS HealthOmics can connect to public and private network resources. AWS HealthOmics is a fully managed bioinformatics service designed to accelerate scientific breakthroughs at scale for clinical diagnostics, drug discovery, and agriculture research customers.
In this post you will discover the benefits and use cases for VPC connected workflows, the steps to configure a VPC endpoint, practical walkthroughs for common scenarios, and how VPC security controls apply to workflow traffic.
The Challenge
Previously, AWS HealthOmics supported a restricted network mode for running workflows. In restricted mode, workflows run in an isolated compute environment with no outbound network access. That means workflows can only reach a predefined set of AWS services and have no ability to connect to the public internet, private repositories, or external data sources during execution.
Running bioinformatics pipelines that require network resources in restricted mode added operational overhead. External dependencies had to be manually pre-staged into S3 before a workflow could run, turning data access into a recurring preparation task. For teams working with frequently updated databases like ClinVar or dbSNP, that meant scheduled data transfers, version tracking across pipeline runs, and pipeline failures when a pre-staged file fell out of date.
The Solution
AWS HealthOmics now supports VPC connected workflows, a new capability that gives workflows outbound network access through your Amazon VPC at runtime. Where restricted mode limited workflows to a fixed set of AWS services, VPC connected workflows route traffic through a VPC endpoint powered by AWS PrivateLink, opening connectivity to AWS services, resources inside your VPC, and the public internet.
A VPC endpoint acts as a private entry point into your network fabric. When a HealthOmics workflow runs in VPC connected mode, the workflow routes all outbound traffic through that endpoint. From there, traffic can reach private resources like internal databases, license servers, or continue to public repositories through a Network Address Translation gateway (NAT gateway) configured in your VPC.
Benefits
VPC connected workflows give your HealthOmics pipelines direct access to the public internet and private repositories at runtime, removing the preparation work that slows workflow deployments. You can pull from live sources like NCBI, Ensembl, and licensed annotation services mid-run, so pipelines always work against current data. That connectivity also simplifies how workflows are built with fewer staging steps and faster time from development to production.
With VPC connected workflows, your genomics pipelines can reach resources that previously inaccessible or required manual workarounds, including:
- Third-party license servers such as Sentieon. Automate license validation and inspect connections with VPC Flow Logs, rather than relying on the HealthOmics service-hosted Sentieon proxy.
- AWS License Manager and use software purchased in the AWS Marketplace directly from your workflows.
- AWS Secrets Manager to retrieve credentials securely at runtime instead of embedding passwords or tokens in your workflow code.
- Resources in your private VPC, such as Amazon DynamoDB tables, Amazon RDS databases, or other internal services.
- S3 Tables to load variant data files for analytics and query thus eliminating the intermediate step of writing to Amazon S3 and building your own data loader.
- S3 buckets across AWS Regions without standing up AWS HealthOmics workflows in a new Region.
- Public datasets at runtime without migrating them into your own storage first.
- External servers, including FTP and SFTP endpoints, for data transfer during workflow tasks.
How VPC connected workflows route traffic
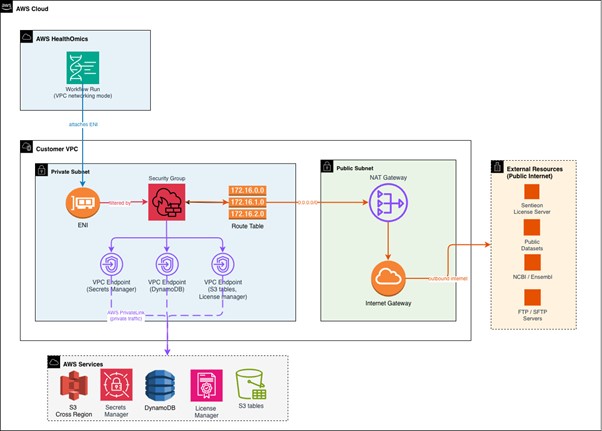
When you enable VPC networking mode on a run, AWS HealthOmics attaches an Elastic Network Interface (ENI) in your private subnet. Outbound traffic from workflow tasks routes through your VPC, passes through the NAT gateway in your public subnet and reaches external resources. For AWS services, the traffic is routed to VPC endpoints. Your security groups and network access control lists (NACL) govern which destinations are reachable. Domain Name Server (DNS) requests are resolved at the configured VPC DNS Resolver.
Figure 1: VPC connected workflows network diagram
Getting Started
Prerequisites
Before you begin, verify your network infrastructure meets these requirements:
You need a VPC with both public and private subnets. If you do not already have one, you can create a new VPC for this purpose. Consider using an AWS CloudFormation template to deploy the required infrastructure (public and private subnets, NAT gateway, and route tables). See the Amazon VPC User Guide for step-by-step instructions.
The private subnets host your HealthOmics workflow elastic network interfaces (ENI), while public subnets contain the NAT gateways that routes outbound traffic to external data sources.
At least one, NAT gateway must exist in a public subnet within your VPC. This gateway enables workflows running in private subnets to reach external resources and public repositories at runtime with a recommended three NAT gateways (one per AZ) for prod environments that require high availability.
Your private subnet route table must direct outbound traffic (0.0.0.0/0) to the NAT gateway. Confirm this routing configuration before proceeding with endpoint setup.
At minimum, your Identity and Access Management principal requires permissions to create VPC endpoints (ec2:CreateVpcEndpoint), describe VPC resources (ec2:DescribeVpcs, ec2:DescribeSubnets, ec2:DescribeSecurityGroups), modify security groups (ec2:AuthorizeSecurityGroupIngress), and configure HealthOmics workflows (omics:CreateWorkflow, omics:StartRun). Review your IAM policies and see the AWS HealthOmics Developer Guide for a full policy reference.). Review your IAM policies and see the AWS HealthOmics Developer Guide for a full policy reference.
For detailed VPC architecture guidance, see Connecting workflows to a VPC in the AWS HealthOmics Developer Guide.
HealthOmics console walkthrough
Connecting a VPC endpoint to HealthOmics is a network configuration change, not a code change. Most teams complete the setup in under an hour.
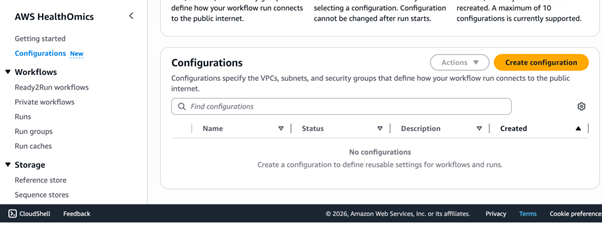
Start by creating a VPC interface endpoint for the HealthOmics service. In the AWS Management Console, navigate to the Configurations menu and choose Create Configuration.
Figure 2: AWS HealthOmics console, configuration menu option
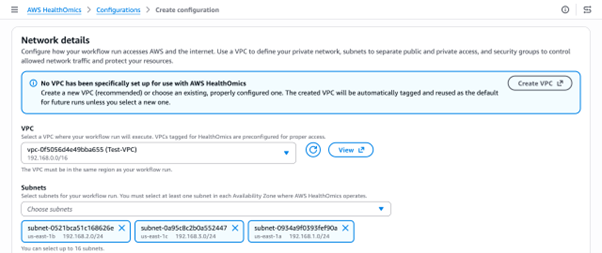
Select a VPC and subnets to associate with the endpoint. A minimum of two subnets is recommended for high availability.
Figure 3: AWS HealthOmics console, associate the VPC and Subnets
Associate a security group with the endpoint to control which external resources a workflow is allowed to reach.
Figure 4: AWS HealthOmics console, associate Security Groups
Best Practice: Validate External URI Connectivity with a Lightweight Test Workflow
Before running production bioinformatics pipelines that depend on external data sources (e.g., NIH datasets, NCBI repositories, third-party license servers, or cross-region S3 buckets), run a minimal connectivity test workflow in VPC networking mode to confirm that runtime data fetch succeeds end-to-end.
Why this matters: AWS HealthOmics runs in RESTRICTED mode by default, which blocks all public internet and cross-region access. Switching to VPC mode routes traffic through your NAT gateway but misconfigured security groups, missing NAT gateway routes, or incorrect IAM permissions will fail at runtime, not at workflow deployment time. A test run catches these issues early.
Start the workflow in VPC Mode
Key enhancements in this command:
- –networking-mode VPC: Enables VPC networking mode for secure, private network access to resources
- –configuration-name my-vpc-config: References your pre-configured VPC settings (subnets, security groups)
For full configuration details, see the AWS HealthOmics Developer Guide.
Use case walkthroughs
The following walkthroughs cover the most common scenarios. Each assumes you have already completed the VPC endpoint setup above.
Connect to a Sentieon license server
If you use Sentieon tools in your genomics workflows, VPC connected workflows let you connect directly to your Sentieon license server instead of relying on the HealthOmics service-hosted proxy.
To set this up, configure your security group to allow outbound traffic to the license server IP address and port. In your workflow task, set the SENTIEON_LICENSE environment variable to point to the license server endpoint. Your workflow tasks validate licenses at runtime through your VPC, and you can monitor these connections using VPC Flow Logs.
Connect to Amazon DynamoDB in a task
VPC connected workflows can read from and write to DynamoDB tables during workflow tasks, which is useful for storing metadata, tracking sample status, or recording pipeline results.
Create a VPC gateway endpoint for DynamoDB
(com.amazonaws.<region>.dynamodb) in the same VPC. Your workflow task can then use the AWS SDK to call GetItem, PutItem, or Query. Ensure the IAM role attached to your workflow run has the required DynamoDB permissions. Because DynamoDB uses a gateway endpoint rather than an interface endpoint, there is no additional hourly charge for the endpoint itself.
Write VCF outputs directly to S3 Tables
With VPC connected workflows, your tasks can write VCF file outputs directly to an S3 Table bucket, removing the need to first write results to Amazon S3 and then run a separate data loader.
Configure your task to target the S3 Tables endpoint in your VPC. Ensure your security group allows outbound HTTPS traffic (port 443) to the S3 Tables service. The IAM role for your workflow run needs s3tables:PutTableData and s3tables:GetTableData permission on the target table. This approach lets downstream analytics tools query your VCF results immediately after a workflow completes. For a sample workflow, reference variant data to S3 table loader in AWS HealthOmics Tutorials.
Access S3 data across AWS Regions
If your reference data or input files live in S3 buckets in a different AWS Region, VPC connected workflows let you access them without replicating the data or deploying HealthOmics workflows in that Region.
Your workflow tasks can make cross-Region S3 API calls through the NAT gateway. Ensure your security group allows outbound HTTPS traffic and that the IAM role has s3:GetObject permission on the remote bucket. Be aware that cross-Region data transfer charges apply. When using VPC networking mode, you are responsible for determining whether it is safe and compliant to transfer or use data across AWS Regions.
Connect to an FTP or SFTP server
Some genomics data sources still use FTP for distribution. VPC connected workflows can reach these servers at runtime.
For FTP, your security group must allow outbound traffic on the control port (typically port 21) and the passive data port range used by the server. For FTP servers on the public internet, traffic routes through your NAT gateway. For servers inside your VPC, ensure the route table and security groups permit direct communication between the workflow subnet and the server subnet.
Where possible, prefer SFTP (port 22) or FTPS (port 990) over plain FTP. These protocols encrypt data in transit, which is important when transferring sensitive genomics data. Adjust your security group rules to allow the appropriate port for your chosen protocol.
For SFTP install your container dependencies. Don’t yum install at runtime in VPC mode. Package managers may fail due to DNS resolution or repo connectivity issues, and set -e will terminate the task immediately. The only required package for SFTP is openssh-clients, which provides the sftp, ssh, and ssh-keygen commands. The base image public.ecr.aws/amazonlinux/amazonlinux:2023 does not include these by default:
openssh-clients also covers SCP transfers and remote SSH commands. For password-based SFTP authentication with a custom AWS Transfer Family identity provider, additionally include sshpass.
If you are using AWS Transfer as your SFTP/FTP server, prefer a Transfer Family VPC endpoint over routing through a NAT gateway. This is distinct from the HealthOmics VPC endpoint, which routes HealthOmics traffic, whereas the Transfer Family VPC endpoint keeps your SFTP connection on the AWS private network, eliminating NAT gateway data processing charges and SSH algorithm compatibility issues that arise when Transfer Family’s service-managed identity provider evaluates connections from a NAT gateway IP.
Use AWS Marketplace software with License Manager
VPC connected workflows let you use software purchased through the AWS Marketplace and managed by AWS License Manager directly in your workflow tasks.
Create a VPC endpoint for the License Manager service (com.amazonaws.<region>.license-manager) in the same VPC. Your workflow task can then call the License Manager API via the AWS SDK to check out a license at the start of a task and check it back in when the task completes. Ensure the IAM role attached to your workflow run includes license-manager:CheckoutLicense and license-manager:CheckinLicense permissions.
Retrieve credentials with Secrets Manager
Instead of embedding passwords or API tokens in your workflow code, you can retrieve them securely at runtime from AWS Secrets Manager.
Create a VPC endpoint for Secrets Manager (com.amazonaws.<region>.secretsmanager) in your VPC. In your workflow task, use the AWS SDK to call GetSecretValue with the secret ARN. Your security group must allow outbound HTTPS traffic (port 443) to the Secrets Manager endpoint. Grant the workflow run IAM role the secretsmanager:GetSecretValue permission for the specific secret ARN.
Security controls
VPC connected workflows are built with multiple layers of security. By default, VPC networking is disabled for all workflow runs. To connect a workflow to external resources, you must explicitly set up a VPC endpoint and enable VPC networking mode on the run. No workflow can reach outside the HealthOmics environment unless you opt in.
Once VPC networking mode is enabled, your existing VPC security controls govern all workflow traffic. Security group rules specify which IP addresses, CIDR ranges, and ports each workflow can reach. Network ACLs provide an additional layer of subnet-level filtering. For domain-based filtering, use Amazon Route 53 Resolver DNS Firewall to control which DNS names your workflows can resolve. VPC Flow Logs capture network activity for audit and compliance review.
Your existing subnet layouts, route tables, and gateway configurations carry over without modification. AWS HealthOmics VPC endpoints are available in every AWS Region where HealthOmics private workflows are supported. Interface VPC endpoints incur hourly and per-GB charges. Review costs before you configure an endpoint on the AWS PrivateLink pricing page.
Call Caching with VPC Connected Workflows
Call caching lets AWS HealthOmics reuse outputs from previous runs when the inputs and task definitions have not changed, reducing compute time and cost across repeated pipeline executions. When workflows connect to external sources through a VPC endpoint, caching behavior requires some additional consideration. Public repositories and annotation services can return different results over time, so tasks that pull from non-deterministic sources should either have caching turned off for the entire run, or be opted out at the task level so their outputs are not stored and reused in future runs.
Nextflow – Turn off caching for individual tasks using the “cache false” directive.
WDL – Disable caching for individual tasks using the “volatile” attribute.
CWL – Control caching behavior for individual tasks using the “WorkReuse” feature.
For additional engine specific guidance reference Engine Specific Caching Features
Next steps
Where you start depends on where you are with AWS HealthOmics today. If you’re new to HealthOmics, begin by visiting the AWS HealthOmics Setup guide to get started, then read the AWS HealthOmics Developer Guide to understand how private workflows handle data access.
If you’re already running workflows on AWS HealthOmics, start by identifying which data sources your workflows fetch manually today. Those are the best candidates to migrate to runtime access. Next, check your existing security group and network ACL rules to confirm they cover the external domains your workflows will reach at runtime. Then open the AWS HealthOmics console and try a VPC endpoint configuration on a non-production workflow first. Finally, send feedback or questions to AWS re:Post for HealthOmics or through your AWS Support contacts.
Try AWS HealthOmics VPC endpoints today. To learn more, visit the AWS HealthOmics product page.