AWS for Industries
Event-Driven Digital Pathology: Governed Whole Slide Image Ingestion to Scalable Inference with Amazon SageMaker
Introduction
Whole-slide images (WSIs) are transforming digital pathology, but working with these massive files across different scanners and systems presents real challenges. This blog post will detail how Genmab, a leading biotech company, built an automated pipeline on AWS that handles whole-slide images from start to finish, cutting analysis time from hours to under 30 minutes per batch and reducing manual work by 80 percent. We will walk through how Genmab achieved this using AWS services and share the key lessons they learned along the way.
Opportunity
Genmab’s digital pathology laboratory scans large tissue image, often several gigabytes each. The team faced a growing challenge managing hundreds of WSIs from multiple sources, each requiring careful tracking, organization, and analysis. While the images, along with metadata were stored in Concentriq, the actual analysis happened on-premises. This setup required significant manual coordination and limited throughput to about 50 slides per day, which could not keep pace with demand. Additionally, the results were stored separately from the WSIs and metadata, making it difficult for teams to get a complete picture and establish a single source of truth.
Solution
To solve this challenge, Genmab built an automated pipeline for processing WSIs. The pipeline runs without manual intervention and uses custom machine learning models in Amazon SageMaker for image analysis. It handles two types of input: complete WSI files and pre-tiled images. For pre-tiled images, the pipeline processes all tiles simultaneously using Amazon SageMaker Batch Transform, which speeds up the analysis. We’ll walk through both approaches, each one suited for specific model and throughput requirements.
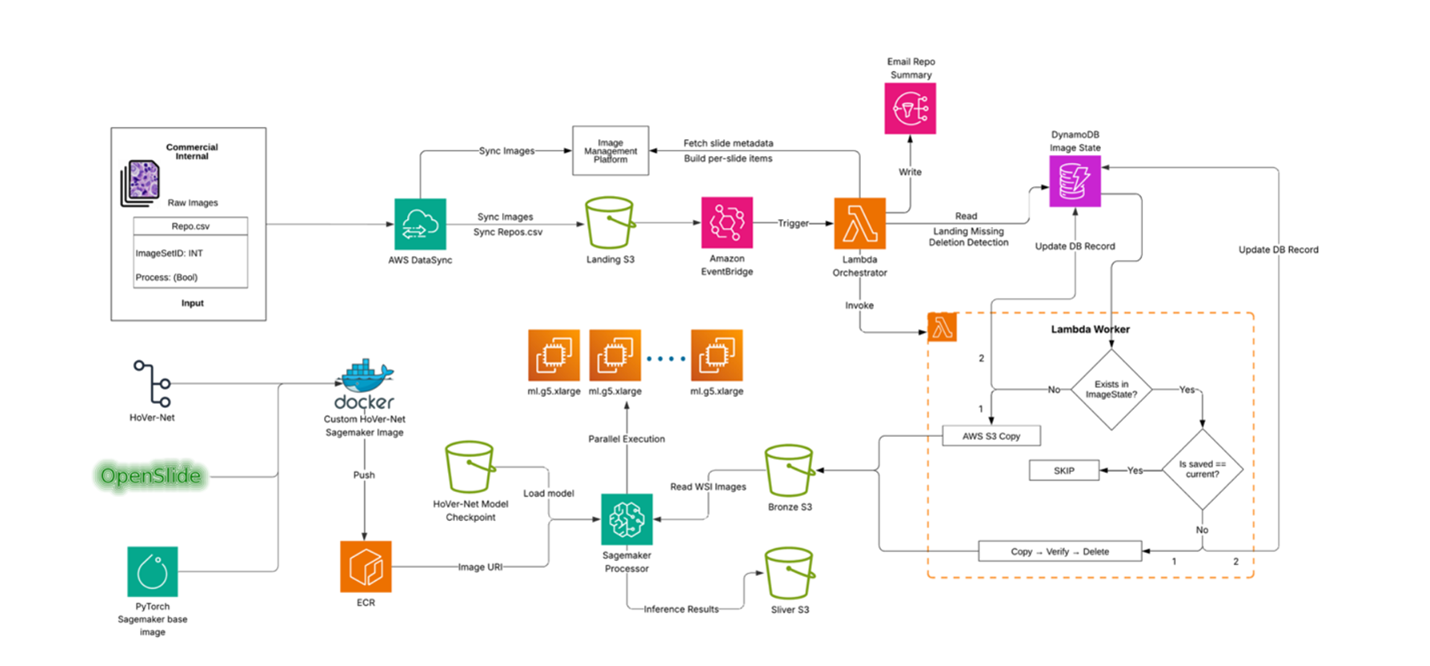
End-to-end Process Flow
Genmab’s digital pathology laboratory continuously writes WSI files to a network drive. The process is triggered when a researcher drops a manifest file (repo.csv) to a network drive, which begins the transfer of files to the Landing Amazon S3 bucket using AWS DataSync.
This S3 event invokes a “coordinator” AWS Lambda function, which retrieves the image metadata from Concentriq based on the manifest and invokes a “worker” AWS Lambda function for each slide.
Each “worker” Lambda copies the large WSI into the Bronze S3 bucket using multipart copy. Any file renaming, moving, or missing data is recorded in Amazon DynamoDB for full lineage and auditing history. The worker Lambda then sends a summary to the researchers via Amazon SNS (Simple Notification Service).
The Bronze S3 event invokes the AI inference in one of two ways:
- Amazon SageMaker Processing: Pulls slides directly from S3 and processes them in parallel using a custom Docker image (PyTorch + OpenSlide + model code). It scales by sharding S3 keys across instances and multiprocessing per node.
- SageMaker Batch Transform: Processes pre-tiled WSIs using packaged PyTorch models, creates a SageMaker Model, and runs Transform jobs with concurrency controls, built-in queuing, scaling, and detailed logs.
Both approaches write results to the Silver S3 bucket.
Containerized Runtime: Docker and Amazon ECR
Diving deeper into containerized runtime, once slides are stored in Bronze, they need to be prepared for AI model processing. This is done by packaging the inference code inside a SageMaker-compatible Docker container. The container starts from a PyTorch base image, installs OpenSlide to read WSI formats, and includes the model code. The entry point script loads the model and runs inference. Because the container is portable, it can run in either SageMaker Processing or Batch Transform with minimal changes.
After building, the container image is pushed to Amazon ECR (Elastic Container Registry), making it easy to version, reuse, and swap in new models. This design keeps the pipeline model-agnostic: Genmab can run HoVer-Net today, ResNet50 tomorrow, or an entirely new model later. This flexibility is achieved simply by pointing SageMaker to a different container image.
Option 1: Scalable Inference with SageMaker Processing
SageMaker Processing is used for models that operate on whole slides and can scale horizontally. SageMaker distributes input slides across multiple GPU instances, while each instance parallelizes work internally across CPU and GPU workers. This means hundreds of gigabyte-scale WSIs can be analyzed in parallel without maintaining clusters or manually scheduling GPUs. Jobs can be tuned with parameters like tile size, chunk size, and worker counts to balance GPU memory with throughput. Amazon CloudWatch provides visibility into performance and allows setting adjustments as data or models change.
Option 2: Scalable Inference with Batch Transform
SageMaker Batch Transform is used for models that work with pre-tiled images. Slides are pre-tiled, background regions are filtered out, and the resulting tiles are uploaded to S3. Batch Transform then processes millions of tiles with built-in queuing and concurrency controls.
To use this approach, a model (such as a vision model) is packaged into a model.tar.gz bundle that includes the model weights and inference script. The package is uploaded to S3, a SageMaker Model is created, and a Transform job is launched.
Batch Transform is well suited for high-throughput offline inference on pre-tiled datasets. It can process large tile datasets efficiently using managed batch jobs without deploying or operating real-time endpoints. Metrics like transform latency and queue time in CloudWatch help tune payload sizes and concurrency.
Overall Performance and Cost Tuning
Scaling WSI inference isn’t just about raw compute—it’s about balancing throughput, cost, and GPU (graphics processing unit) constraints, such as accelerator memory limits and, in some cases, account quotas. Here are a few lessons learned along the way:
Tile and chunk sizes matter — Choose tile and chunk sizes carefully. Start small and grow until GPU memory is fully utilized.
Avoid oversubscription — Set thread counts carefully to prevent CPUs from thrashing.
Horizontal scale beats vertical scale — Increasing instance count often gives better throughput per dollar than using larger instances.
Cache smartly — Caching tiles on ephemeral storage avoids costly re-reads of entire WSIs.
These practices allowed Genmab to run thousands of gigabyte-scale slides reliably while staying cost-aware.
Security, Networking, and Governance
Governance is central to this pipeline. Every step follows least-privilege AWS Identity and Access Management (IAM) principles. Lambda functions, SageMaker jobs, and DynamoDB tables only access the resources they need.
Data is encrypted at rest and in transit, and all jobs run inside VPC subnets with S3 interface endpoints for controlled networking.
For auditability, manifests are maintained in S3, detailed lineage is tracked in DynamoDB, and structured logs are stored in CloudWatch. Together, these artifacts create an audit trail that ties any output back to its exact input and parameters. CloudWatch Alarms detect errors, stale control files, or stuck objects in the Landing bucket.
Troubleshooting
To handle common issues, the system detects, retries, and surfaces errors clearly:
Missing slides — If a slide is missing from the Landing bucket, the worker marks it as missing and automatically ingests it later if it reappears.
Failed renames — If a rename fails, the worker retries with a safe fallback.
Duplicate files — If duplicates appear, they can be resolved upstream in the image management system and retried.
Slow Batch Transform jobs — If jobs slow down, adjusting tile payload size or concurrency usually resolves the issue.
These safeguards mean operators don’t need to manually monitor the system—errors are logged, retried, and surfaced clearly through Amazon SNS and CloudWatch.
Conclusion
This solution removes the complexity of working with large pathology images at scale. Lab users don’t need to learn AWS or manage files manually—a simple CSV starts the process. The system moves images into the right location, tracks every change, and reports back automatically.
Genmab’s automated pipeline transformed their digital pathology workflow, reducing manual coordination time by 80 percent and increasing processing capacity from tens to hundreds of slides daily. What previously took hours and was limited to approximately 50 slides per day now runs continuously with near-real-time ingestion and AI inference.
The pipeline is secure, auditable, and built on managed AWS services, giving Genmab a scalable foundation that unifies data governance and large-scale computation. This accelerates analysis cycles and improves reliability across their digital pathology workflows.
To learn more, see Amazon Batch Transform Documentation and AWS SageMaker AI Blogs.