AWS HPC Blog
Category: *Post Types
Bayesian ML Models at Scale with AWS Batch
Ampersand is a data-driven TV advertising technology company that provides aggregated TV audience impression insights and planning on 42 million households, in every media market, across more than 165 networks and apps and in all dayparts (broadcast day segments). The Ampersand Data Science team estimated that building their statistical models would require up to 600,000 physical CPU hours to run, which would not be feasible without using a massively parallel and large-scale architecture in the cloud. AWS Batch enabled Ampersand to compress their time of computation over 500x through massive scaling while optimizing their costs using Amazon EC2 Spot. In this blog post, we will provide an overview of how Ampersand built their TV audience impressions (“impressions”) models at scale on AWS, review the architecture they have been using, and discuss optimizations they conducted to run their workload efficiently on AWS Batch.
Introducing the Spack Rolling Binary Cache hosted on AWS
Today we’re excited to announce the availability of a new public Spack Binary Cache. In a collaboration, between AWS, E4S, Kitware, and the Lawrence Livermore National Laboratory (LLNL), Spack users now have access to a public build cache hosted on Amazon S3. The use of this Binary Cache will result in up to 20x faster install times for common Spack packages.
Encoding workflow dependencies in AWS Batch
This post covers the different ways you can encode a dependency between basic and array jobs in AWS Batch. We also cover why you may want to encode dependencies outside of Batch altogether using a workflow system like AWS Step Functions or Apache Airflow.
Benchmarking NVIDIA Clara Parabricks Somatic Variant Calling Pipeline on AWS
Somatic variants are genetic alterations which are not inherited but acquired during one’s lifespan, for example those that are present in cancer tumors. In this post, we will demonstrate how to perform somatic variant calling from matched tumor and normal genome sequence data, as well as tumor-only whole genome and whole exome datasets using an NVIDIA GPU-accelerated Parabricks pipeline, and compare the results with baseline CPU-based workflows.
AI-based drug discovery with Atomwise and WEKA Data Platform
Drug discovery is an expensive proposition, with a $2.6 billion cost over 10 years and just a 12% success rate. AI promises to significantly improve the success rate by finding small molecule hits for undruggable targets. On the forefront of using AI in drug discovery is Atomwise, with its AtomNet® platform. In this blog, we will lay out the challenges of the drug discovery process, and show how AI/ML startups are solving these challenges using solutions from Atomwise, AWS, and WEKA.
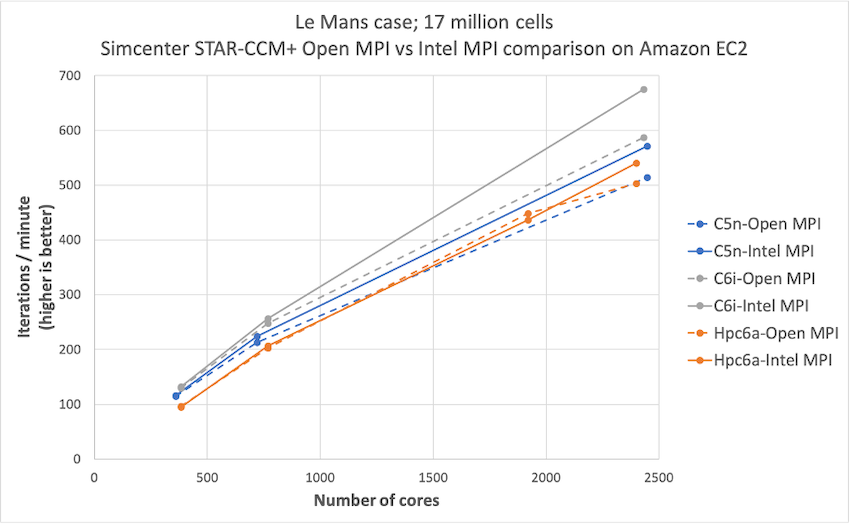
Simcenter STAR-CCM+ price-performance on AWS
Organizations such as Amazon Prime Air and Joby Aviation use Simcenter STAR-CCM+ for running CFD simulations on AWS so they can reduce product manufacturing cycles and achieve faster times to market. In this post today, we describe the performance and price analysis of running Computational Fluid Dynamics (CFD) simulations using Siemens SimcenterTM STAR-CCM+TM software on AWS HPC clusters.
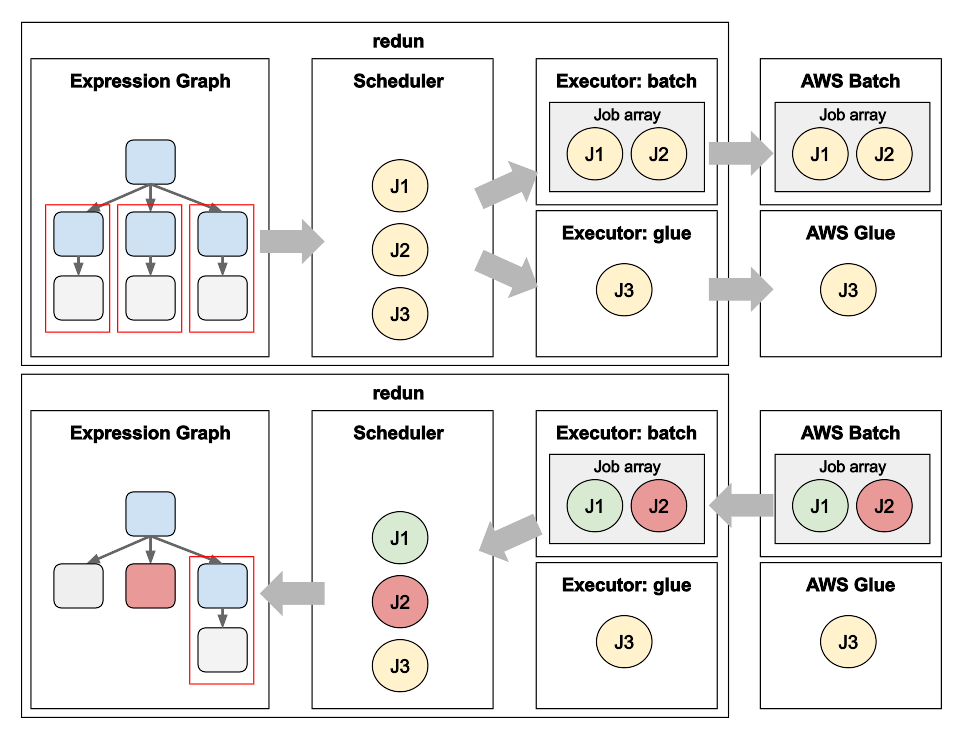
Data Science workflows at insitro: how redun uses the advanced service features from AWS Batch and AWS Glue
Matt Rasmussen, VP of Software Engineering at insitro, expands on his first post on redun, insitro’s data science tool for bioinformatics, to describe how redun makes use of advanced AWS features. Specifically, Matt describes how AWS Batch’s Array Jobs is used to support workflows with large fan-out, and how AWS Glue’s DynamicFrame is used to run computationally heterogenous workflows with different back-end needs such as Spark, all in the same workflow definition.
Data Science workflows at insitro: using redun on AWS Batch
Matt Rasmussen, VP of Software Engineering at insitro describes their recently released, open-source data science framework, redun, which allows data scientists to define complex scientific workflows that scale from their laptop to large-scale distributed runs on serverless platforms like AWS Batch and AWS Glue. I this post, Matt shows how redun lends itself to Bioinformatics workflows which typically involve wrapping Unix-based programs that require file staging to and from object storage. In the next blog post, Matt describes how redun scales to large and heterogenous workflows by leveraging AWS Batch features such as Array Jobs and AWS Glue features such as Glue DynamicFrame.
Creating a digital map of COVID-19 virus for discovery of new treatment compounds
Quantum physics and high-performance computing have slashed research times for a consortium of researchers led by Qubit Pharmaceuticals. This post describes the discovery of chemical substances that may lead to new COVID-19 treatments in only six months using cloud technology.
Choosing between AWS Batch or AWS ParallelCluster for your HPC Workloads
It’s an understatement that AWS has a lot of services (more than 200 at the time of this post!). We’re usually the first to point out that there’s more than one way to solve a problem. HPC is no different in this regard, because we offer a choice: customers can run their HPC workloads using AWS […]