AWS Big Data Blog
Accelerate large-scale data migration validation using PyDeequ
March 2023: You can now use AWS Glue Data Quality to measure and manage the quality of your data. AWS Glue Data Quality is built on DeeQu and it offers a simplified user experience for customers who want to this open-source package. Refer to the blog and documentation for additional details.
Many enterprises are migrating their on-premises data stores to the AWS Cloud. During data migration, a key requirement is to validate all the data that has been moved from on premises to the cloud. This data validation is a critical step and if not done correctly, may result in the failure of the entire project. However, developing custom solutions to determine migration accuracy by comparing the data between the source and target can often be time-consuming.
In this post, we walk through a step-by-step process to validate large datasets after migration using PyDeequ. PyDeequ is an open-source Python wrapper over Deequ (an open-source tool developed and used at Amazon). Deequ is written in Scala, whereas PyDeequ allows you to use its data quality and testing capabilities from Python and PySpark.
Prerequisites
Before getting started, make sure you have the following prerequisites:
- An AWS account with access to AWS services
- An Amazon Virtual Private Cloud (Amazon VPC) with a public subnet
- An Amazon Elastic Compute Cloud (Amazon EC2) key pair
- An AWS Identity and Access Management (IAM) policy for AWS Secrets Manager permissions
- A SQL editor to connect to the source database
- IAM roles
EMR_DefaultRoleandEMR_EC2_DefaultRoleavailable in your account
Solution overview
This solution uses the following services:
- Amazon RDS for My SQL as the database engine for the source database.
- Amazon Simple Storage Service (Amazon S3) or Hadoop Distributed File System (HDFS) as the target.
- Amazon EMR to run the PySpark script. We use PyDeequ to validate data between MySQL and the corresponding Parquet files present in the target.
- AWS Glue to catalog the technical table, which stores the result of the PyDeequ job.
- Amazon Athena to query the output table to verify the results.
We use profilers, which is one of the metrics computation components of PyDeequ. We use this to analyze each column in the given dataset to calculate statistics like completeness, approximate distinct values, and data types.
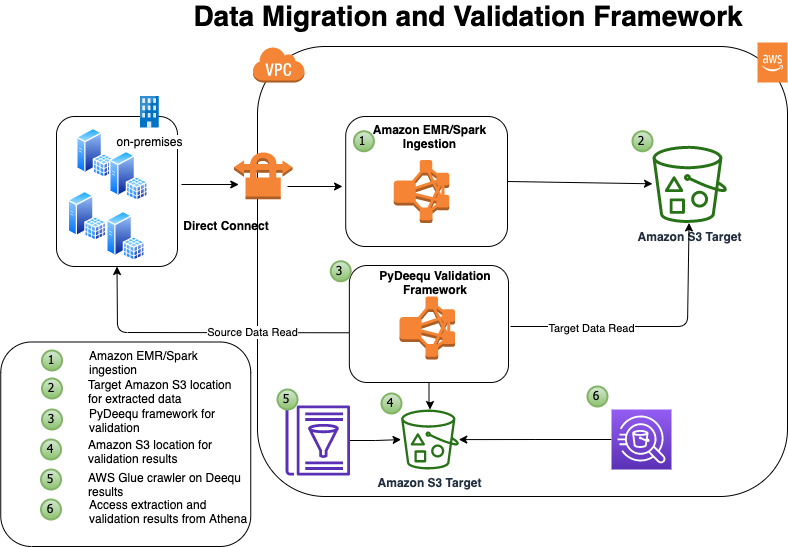
The following diagram illustrates the solution architecture.
In this example, you have four tables in your on-premises database that you want to migrate: tbl_books, tbl_sales, tbl_venue, and tbl_category.
Deploy the solution
To make it easy for you to get started, we created an AWS CloudFormation template that automatically configures and deploys the solution for you.
The CloudFormation stack performs the following actions:
- Launches and configures Amazon RDS for MySQL as a source database
- Launches Secrets Manager for storing the credentials for accessing the source database
- Launches an EMR cluster, creates and loads the database and tables on the source database, and imports the open-source library for PyDeequ at the EMR primary node
- Runs the Spark ingestion process from Amazon EMR, connecting to the source database, and extracts data to Amazon S3 in Parquet format
To deploy the solution, complete the following steps:
The template is launched in the US East (N. Virginia) Region by default.
- On the Select Template page, keep the default URL for the CloudFormation template, then choose Next.
- On the Specify Details page, provide values for the parameters that require input (see the following screenshot).
- Choose Next.
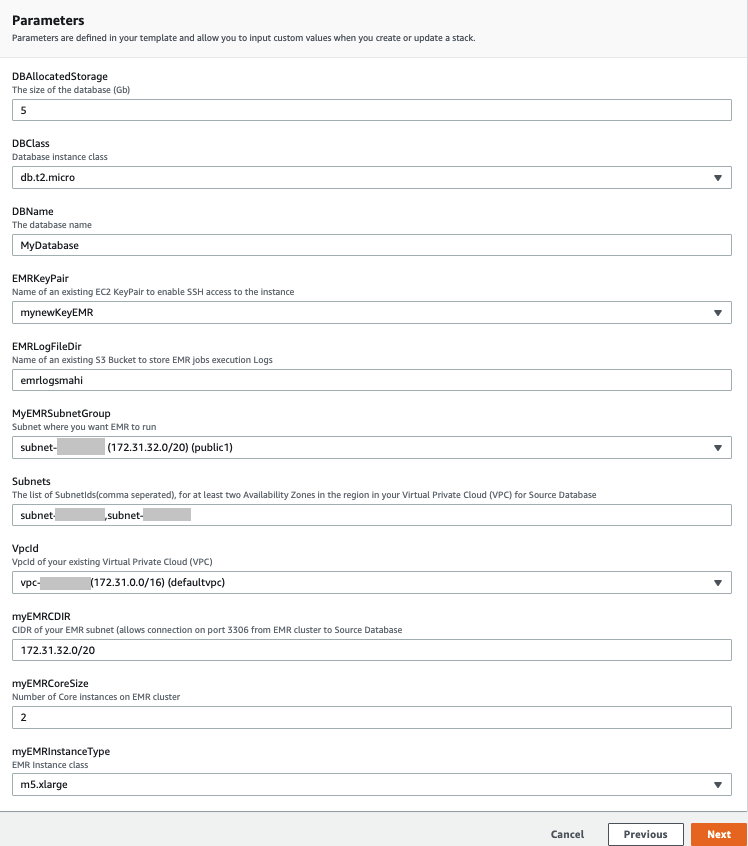
- Choose Next again.
- On the Review page, select I acknowledge that AWS CloudFormation might create IAM resources with custom names.
- Choose Create stack.
You can view the stack outputs on the AWS Management Console or by using the following AWS Command Line Interface (AWS CLI) command:
It takes approximately 20–30 minutes for the deployment to complete. When the stack is complete, you should see the resources in the following table launched and available in your account.
| Resource Name | Functionality |
| DQBlogBucket | The S3 bucket that stores the migration accuracy results for the AWS Glue Data Catalog table |
| EMRCluster | The EMR cluster to run the PyDeequ validation process |
| SecretRDSInstanceAttachment | Secrets Manager for securely accessing the source database |
| SourceDBRds | The source database (Amazon RDS) |
When the EMR cluster is launched, it runs the following steps as part of the post-cluster launch:
- DQsetupStep – Installs the Deequ JAR file and MySQL connector. This step also installs Boto3 and the PyDeequ library. It also downloads sample data files to use in the next step.
- SparkDBLoad – Runs the initial data load to the MySQL database or table. This step creates the test environment that we use for data validation purposes. When this step is complete, we have four tables with data on MySQL and respective data files in Parquet format on HDFS in Amazon EMR.
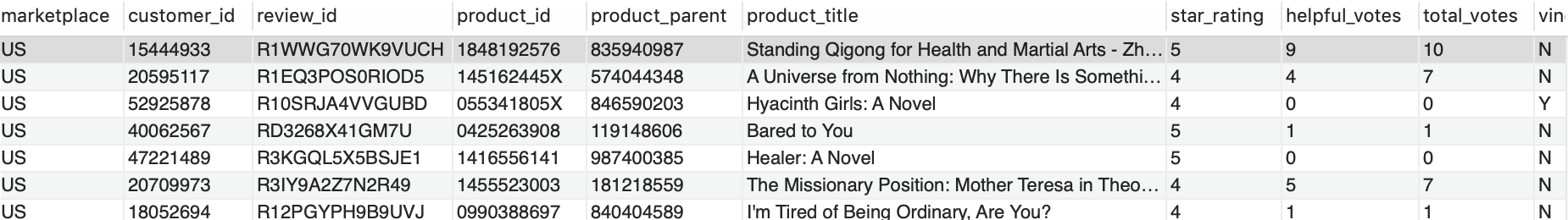
When the Amazon EMR step SparkDBLoad is complete, we verify the data records in the source tables. You can connect to the source database using your preferred SQL editor. For more details, see Connecting to a DB instance running the MySQL database engine.
The following screenshot is a preview of sample data from the source table MyDatabase.tbl_books.
Validate data with PyDeequ
Now the test environment is ready and we can perform data validation using PyDeequ.
- Use Secure Shell (SSH) to connect to the primary node.
- Run the following Spark command, which performs data validation and persists the results to an AWS Glue Data Catalog table (
db_deequ.db_migration_validation_result):
The script and JAR file are already available on your primary node if you used the CloudFormation template. The PySpark script computes PyDeequ metrics on the source MySQL table data and target Parquet files in Amazon S3. The metrics currently calculated as part of this example are as follows:
- Completeness to measure fraction of not null values in a column
- Approximate number of distinct values
- Data type of column
If required, we can compute more metrics for each column. To see the complete list of supported metrics, see the PyDeequ package on GitHub.
The output metrics from the source and target are then compared using a PySpark DataFrame.
When that step is complete, the PySpark script creates the AWS Glue table db_deequ.db_migration_validation_result in your account, and you can query this table from Athena to verify the migration accuracy.
Verify data validation results with Athena
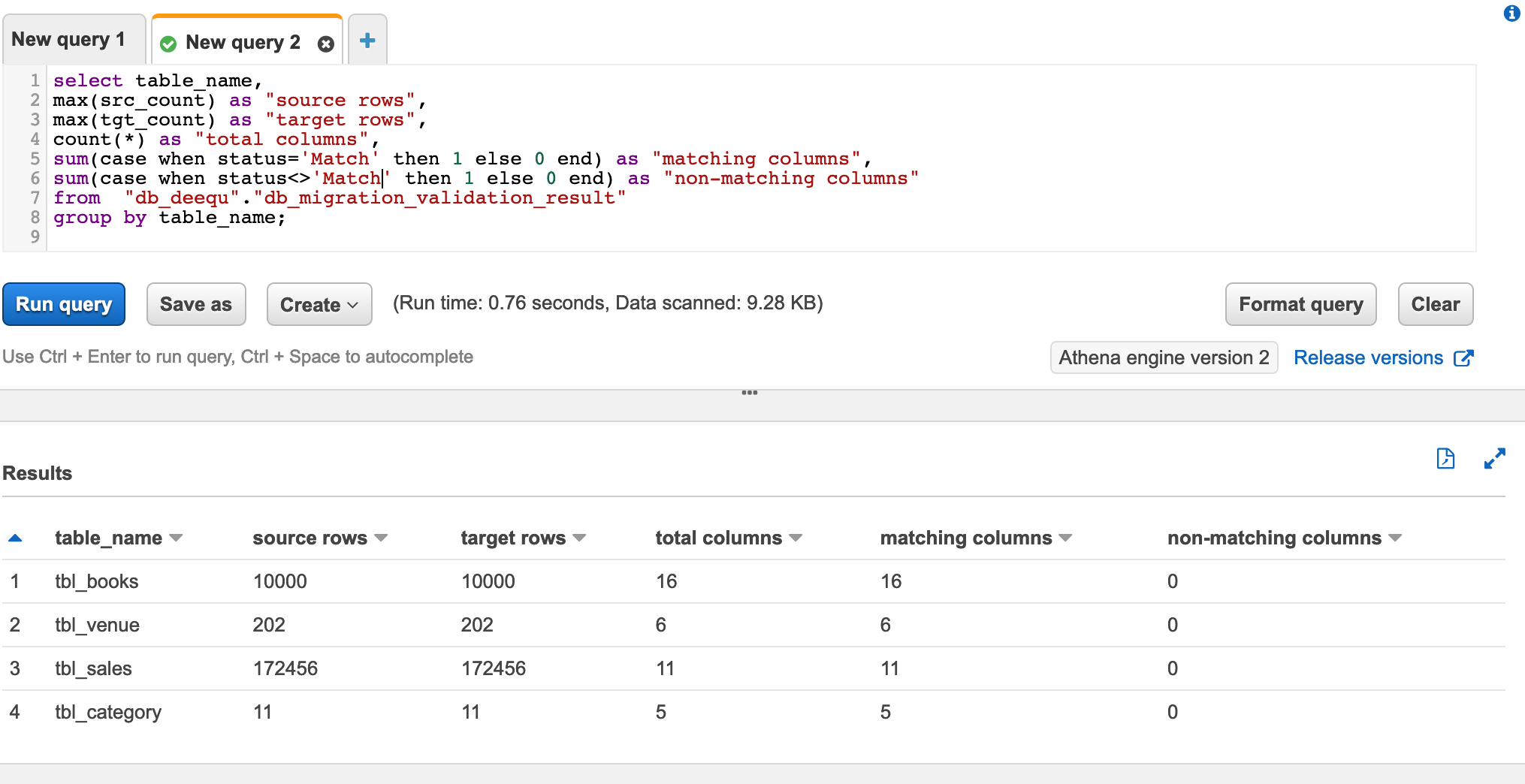
You can use Athena to check the overall data validation summary of all the tables. The following query shows you the aggregated data output. It lists all the tables you validated using PyDeequ and how many columns match between the source and target.
The following screenshot shows our results.
Because all your columns match, you can have high confidence that the data has been exported correctly.
You can also check the data validation report for any table. The following query gives detailed information about any specific table metrics captured as part of PyDeequ validation:
The following screenshot shows the query results. The last column status is the validation result for the columns in the table.
Clean up
To avoid incurring additional charges, complete the following steps to clean up your resources when you’re done with the solution:
- Delete the AWS Glue database and table
db_deequ.db_migration_validation_result. - Delete the prefixes and objects you created from the bucket
dqblogbucket-${AWS::AccountId}. - Delete the CloudFormation stack, which removes your additional resources.
Customize the solution
The solution consists of two parts:
- Data extraction from the source database
- Data validation using PyDeequ
In this section, we discuss ways to customize the solution based on your needs.
Data extraction from the source database
Depending on your data volume, there are multiple ways of extracting data from on-premises database sources to AWS. One recommended service is AWS Data Migration Service (AWS DMS). You can also use AWS Glue, Spark on Amazon EMR, and other services.
In this post, we use PySpark to connect to the source database using a JDBC connection and extract the data into HDFS using an EMR cluster.
The primary reason is that we’re already using Amazon EMR for PyDeequ, and we can use the same EMR cluster for data extraction.
In the CloudFormation template, the Amazon EMR step SparkDBLoad runs the PySpark script blogstep3.py. This PySpark script uses Secrets Manager and a Spark JDBC connection to extract data from the source to the target.
Data validation using PyDeequ
In this post, we use ColumnProfilerRunner from the pydeequ.profiles package for metrics computation. The source data is from the database using a JDBC connection, and the target data is from data files in HDFS and Amazon S3.
To create a DataFrame with metrics information for the source data, use the following code:
Similarly, the metrics is computed for the target (the data file).
You can create a temporary view from the DataFrame to use in the next step for metrics comparison.
After we have both the source (vw_Source) and target (vw_Target) available, we use the following query in Spark to generate the output result:
The generated result is stored in the db_deequ.db_migration_validation_result table in the Data Catalog.
If you used the CloudFormation template, the entire PyDeequ code used in this post is available at the path /home/hadoop/pydeequ_validation.py in the EMR cluster.
You can modify the script to include or exclude tables as per your requirements.
Conclusion
This post showed you how you can use PyDeequ to accelerate the post-migration data validation process. PyDeequ helps you calculate metrics at the column level. You can also use more PyDeequ components like constraint verification to build a custom data validation framework.
For more use cases on Deequ, check out the following:
- Building a serverless data quality and analysis framework with Deequ and AWS Glue
- Build an automatic data profiling and reporting solution with Amazon EMR, AWS Glue, and Amazon QuickSight
- Testing data quality at scale with PyDeequ
About the Authors
 Mahendar Gajula is a Sr. Data Architect at AWS. He works with AWS customers in their journey to the cloud with a focus on Big data, Data Lakes, Data warehouse and AI/ML projects. In his spare time, he enjoys playing tennis and spending time with his family.
Mahendar Gajula is a Sr. Data Architect at AWS. He works with AWS customers in their journey to the cloud with a focus on Big data, Data Lakes, Data warehouse and AI/ML projects. In his spare time, he enjoys playing tennis and spending time with his family.
 Nitin Srivastava is a Data & Analytics consultant at Amazon Web Services. He has more than a decade of datawarehouse experience along with designing and implementing large scale Big Data and Analytics solutions. He works with customers to deliver the next generation big data analytics platform using AWS technologies.
Nitin Srivastava is a Data & Analytics consultant at Amazon Web Services. He has more than a decade of datawarehouse experience along with designing and implementing large scale Big Data and Analytics solutions. He works with customers to deliver the next generation big data analytics platform using AWS technologies.