AWS for Industries
Biotech startup builds agile drug discovery pipeline with scalable compute
Blog guest authored by Andreas Wilm, Director of Computational Biology and Lorenz Gerber, Associate Director of Software Engineering, at ImmunoScape.
Discovering next-gen TCR cell therapies can involve processing and analyzing hundreds of multi-omics patient samples regularly. Leveraging the cloud, biotech startup ImmunoScape has been able to analyze more than 20 trillion data points (generating genomics data for more than one million single T-cells) since innovating their new discovery pipeline.
In little over a week, one of their engineers was able to deploy a high-throughput production analysis pipeline, which otherwise would have taken months with traditional infrastructure procurement.
ImmunoScape uses their tried-and-tested platforms for discovering and developing TCR cell therapies, including those for cancer. Their platform analyzes biomedical big data in a high-throughput fashion to provide insights for immunologists. Being a startup in this highly complex field, they have to smartly meet these requirements:
- Access to high-capacity and elastically scaling storage and compute solutions
- Affordable infrastructure solution without requiring upfront capital
- Infrastructure which does not require dedicated specialist staff to be hired
- Instantly available tailored compute environment for data analysis
- Multi-region infrastructure to cater to the Singapore HQ and their branch in San Diego with a consistent experience
Let’s dive deeper into the journey in finding their solution on Amazon Web Services (AWS).
The batch analysis solution
Early 2021, ImmunoScape added T-cell profiling to its sample analytics stack, a single cell genomics technique that produces data. This analysis requires heavy computational demands in terms of CPUs, RAM and input/output. Usually this analysis is run on a small-scale high-performance computing (HPC) cluster.
The associated data comes in batches for immediate analyses. In between batches, the high-throughput compute infrastructure is not needed. Hence the appropriate solution was to implement an analysis pipeline leveraging the elastic and automatic scaling capabilities of AWS Batch.
AWS Batch automatically spins up resources as needed and then automatically scales them down again after the analysis is completed. Customers are charged for compute capacity only when the resources are running, as part of the cloud’s pay-as-you-go model.
The analysis pipeline was implemented in a workflow manager widely used in Computational Biology called Nextflow. Nextflow, originally an academic product, is developed under an open-source license by Seqera Labs, an AWS partner. More importantly, it has support for multiple HPC schedulers as well as AWS Batch built-in.
It took one engineer little over a week to deploy a production version in the cloud, which has in the meantime generated linked multi-omics data for more than one million single T-cells. The alternative would have been to procure an on-site HPC-like cluster which could have taken months.
In addition, an on-site cluster is of fixed capacity and is therefore by definition either over-or under-provisioned for spiky workloads, as is typical for scientific analysis. On the cloud however, compute resources elastically and automatically scale up and down as needed.
Figure 1 shows an example of ImmunoScape’s compute workloads in the cloud ranging from minutes to hours. This requires the compute infrastructure to be elastic and capable of processing jobs concurrently.
Figure 1: ImmunoScape’s compute workloads
“Scientific analysis workloads have notoriously spiky compute demands. Using the cloud’s automatically scaling compute capabilities allow us to analyze data the moment it arrives and to generate timely insights.” ―Andreas Wilm, Director of Computational Biology at ImmunoScape.
With the cloud, ImmunoScape pays only for what they use, whereas an on-site deployment typically has fixed cost and comes with ongoing maintenance costs, whether it’s used at capacity or not.
This analysis pipeline was integrated into ImmunoScape’s web platform Cytographer, which allows scientists to operate such pipelines through a unified web-frontend. Next, we look at how it was built on AWS.
ImmunoScape’s analysis platform (Cytographer)
Cytographer is ImmunoScape’s online analysis and data platform. It offers their scientists data processing and analytics capabilities to discover immunological insights from their data, which leads to the discovery of new TCR-based drugs. Figure 2 shows a screenshot of the main dashboard, with different panels for different analysis, while Figure 3 shows a typical analysis result for high-dimensional data.
Figure 2: Cytographer user interface
Figure 3: Cytographer analysis example
Cytographer enforces versioning of data analytical pipelines and storage of parameters along with analyzed datasets to guarantee reproducibility, which is crucial for scientific work. This was achieved by high modularization and loose coupling of the various processing steps using AWS Batch and various AWS container management services, as well as DevOps principles. These have allowed agile development and deployment without having to focus on the undifferentiated heavy lifting of managing lower level infrastructure.
As a result, integrating new functionality and making it available to their immunologists can be done with minimal effort. Through this system, their scientists in Singapore and San Diego have browser-based access to sophisticated high-throughput analysis capabilities.
“Cytographer provides us with a standardized and consistent method for data analysis and enables tracking and reviewing of data processing at any point in time during the analysis pipeline. It is essential for us that our scientists can securely access and work on the same data independent of their physical location and a deployment of Cytographer on AWS makes this seamlessly possible.” ―says Lorenz Gerber, Associate Director Software Engineering at ImmunoScape.
The architecture
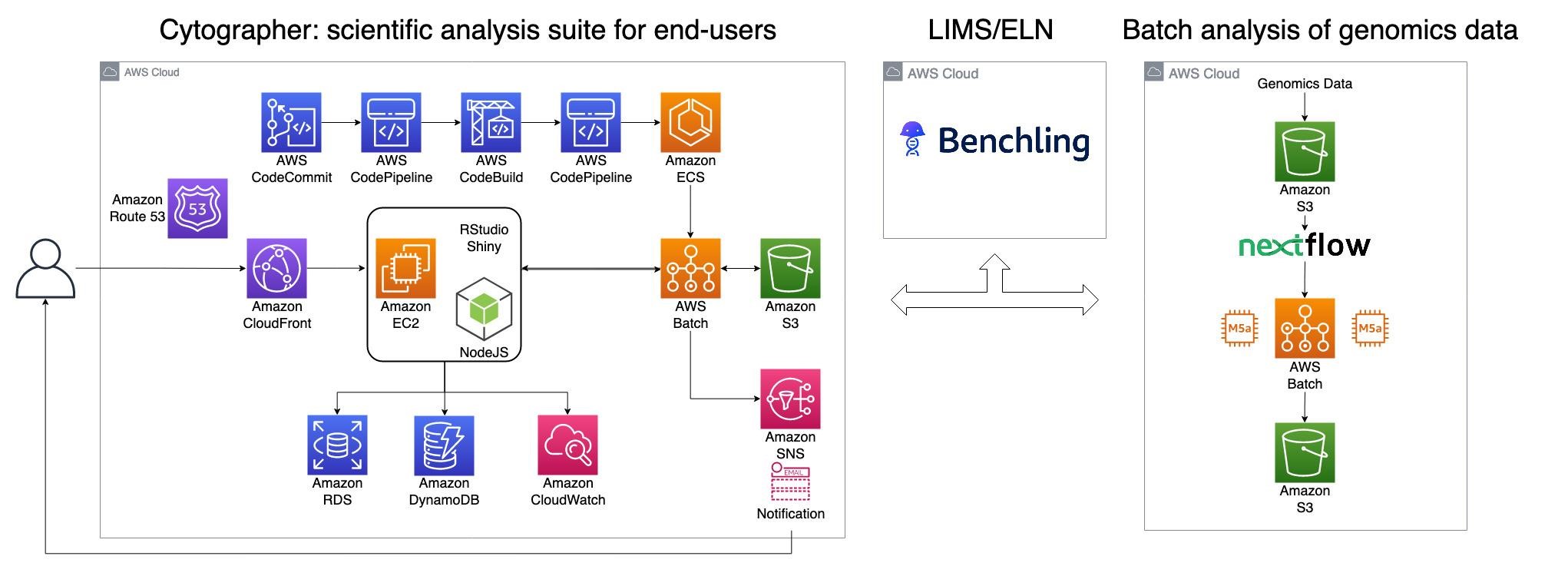
Cytographer leverages multiple components in the cloud. Figure 4 portrays the components used and their interconnections.
Figure 4: Architecture of Cytographer
The user-facing part is a three-tier web application augmented with modular interactive Shiny dashboards served through Shinyproxy. The frontend is served globally through Amazon CloudFront (CloudFront) as a content delivery network (CDN) while analytical data is kept in specific geographic regions for quick and compliant data access and storage.
The heavy data processing runs on AWS Batch. All application and analytical code is DevOps-managed using AWS CodeBuild to build a Docker image for each deployment and AWS CodePipeline as the continuous integration tool. Metadata, like sample information, is stored through Benchling, an AWS hosted Laboratory Information Management System (LIMS) that also serves as an Electronic Lab Notebook (ELN).
Genomics data is processed with workflow manager Nextflow, utilizing AWS Batch as its execution engine. Amazon Simple Storage Service (Amazon S3) is used as a data lake. Amazon S3 is a scalable object storage service designed for 99.999999999% (11 9s) of data durability. It has been used by a number of genomics workloads, including those in these case studies by Neumora and Murdoch University.
Having the scientists located both in Singapore HQ and in San Diego requires a multi-region setup which is addressed by the following strategies:
- Using CloudFront as CDN to provide consistent, low latency, user experience when accessing the UI from both far apart locations.
- Using the same configuration-based backend technologies, AWS Batch and Amazon S3, making replicating to the second region straightforward.
These allow their scientists in both locations to have consistent data upload, processing and analytics capabilities.
DevOps and versioning of pipeline containers are centrally managed in the Singapore region and then automatically replicated to the US West (N. California) region.
Key takeaways
Using the cloud, ImmunoScape, a startup, was able to develop and deploy its new production analysis environment in less than a month. This has, in the meantime, generated genomics data alone for more than one million single T-cells.
The agility provided by AWS has allowed ImmunoScape to start the experiment and iterate quickly without having to worry about infrastructure upfront capital and capacity commitment. Procuring and deploying local hardware would have meant working with fixed capacity and limiting their options, while cloud infrastructure can be contracted and is automatically updated.
ImmunoScape also provides a consistent user experience for scientists in Singapore and San Diego by leveraging multi-regions on AWS. Now, ImmunoScape continues scaling up its analytics services to generate immunological insights to ultimately develop next-gen TCR cell therapies.
“ImmunoScape’s deep immunomics platform generates a plethora of biological data that need to be processed, analyzed and mined in order to advance the development of next generation TCR therapies. AWS provides an ideal environment for us to cope with data complexity while retaining flexibility and scalability—from the early stages of sample screening to the discovery of innovative TCRs for therapeutic development.” says Michael Fehlings, Co-founder and VP for Operations and Technology Development at ImmunoScape.
Further Reading:
- A self-pace workshop on Nextflow with AWS Batch
- The landing pages for Genomics on AWS and HCLS on AWS
- A blog post on detailed AWS Batch configuration for genomics workload
- Apply for AWS Activate credits for startups and reach out to the AWS team
More about ImmunoScape
ImmunoScape is a pre-clinical biotechnology company focused on the discovery and development of next-generation TCR cell therapies in the field of oncology. The company’s proprietary Deep Immunomics technology and machine learning platforms enable highly sensitive, large-scale mining and immune profiling of T cells in cancer patient samples to identify novel, therapeutically relevant TCRs across multiple types of solid tumors. ImmunoScape has multiple discovery programs ongoing and will be progressing towards IND-enabling studies and entry into the clinic. For more information, please visit https://immunoscape.com/.
____________
 Lorenz Gerber, PhD: Lorenz is ImmunoScape’s Associate Director of Software Engineering. With his team, he is taking care of building and maintaining a reliable data processing and storage infrastructure. Lorenz has over 10 years of experience from work in scientific computing and software engineering at various institutes and companies in Sweden, Germany, Japan and Singapore. Most assignments circled around processing and analysis of large datasets from mass spectrometry based high-throughput analytical platforms. Before joining ImmunoScape, he was developing scalable computational workflows on hybrid compute platforms at the Genome Institute of Singapore. He holds a PhD in Biology from the University of Umeå.
Lorenz Gerber, PhD: Lorenz is ImmunoScape’s Associate Director of Software Engineering. With his team, he is taking care of building and maintaining a reliable data processing and storage infrastructure. Lorenz has over 10 years of experience from work in scientific computing and software engineering at various institutes and companies in Sweden, Germany, Japan and Singapore. Most assignments circled around processing and analysis of large datasets from mass spectrometry based high-throughput analytical platforms. Before joining ImmunoScape, he was developing scalable computational workflows on hybrid compute platforms at the Genome Institute of Singapore. He holds a PhD in Biology from the University of Umeå.
____________
 Andreas Wilm, PhD: Andreas is ImmunoScape’s Director of Computational Biology, where his team is responsible for big-data analytics and machine learning. He has worked at the intersection of technology, biology, and computer science for 15 years across research institutes in Germany, Ireland, and Singapore. Immediately prior to joining ImmunoScape, Andreas served as a Cloud solution architect and Data & AI subject matter expert on Microsoft’s Worldwide Public Sector team. In one of his prior roles as Bioinformatics core team lead at the Genome Institute of Singapore, he and his team were responsible for developing scalable computational workflows for analyzing genomics big data on hybrid compute platforms. He holds a Ph.D. in Biology from the University of Duesseldorf.
Andreas Wilm, PhD: Andreas is ImmunoScape’s Director of Computational Biology, where his team is responsible for big-data analytics and machine learning. He has worked at the intersection of technology, biology, and computer science for 15 years across research institutes in Germany, Ireland, and Singapore. Immediately prior to joining ImmunoScape, Andreas served as a Cloud solution architect and Data & AI subject matter expert on Microsoft’s Worldwide Public Sector team. In one of his prior roles as Bioinformatics core team lead at the Genome Institute of Singapore, he and his team were responsible for developing scalable computational workflows for analyzing genomics big data on hybrid compute platforms. He holds a Ph.D. in Biology from the University of Duesseldorf.