AWS Storage Blog
A gene-editing prediction engine with iterative learning cycles built on AWS
NRGene develops cutting-edge genomic analytics products that are reshaping agriculture worldwide. Among our customers are some of the biggest and most sophisticated companies in seed-development, food and beverages, paper, rubber, cannabis, and more.
In the middle of 2020, NRGene joined a consortium of companies and academic institutions to build the best-in-class gene-editing prediction platform to date – targeting both clinical (medical) and agricultural genome editing usages. The consortium, named CRISPR-IL, is funded by the Israel Innovation Authority, and includes members from biotech and agro-tech companies, in addition to academic research centers and medical institutions.
CRISPR technology is a novel form of gene editing that enables precise modifications to organisms’ genetic makeup at the genetic level, with unprecedented precision and cost efficiency. The importance of CRISPR – for which its co-discoverers received the 2020 Nobel Prize in Chemistry – lies in its potential in clinical and agricultural use, such as: to facilitate new treatments of human illnesses, and to improve agricultural produce rapidly and safely, especially amid a world of growing populations and dramatic climate changes. The consortium’s work centers around CRISPR editing effectiveness and specificity, including both the prediction and the detection of “editing events” – part of a broader global research around CRISPR technologies that includes development of new enzymes, delivery systems, usage cases, high-throughput application and much more.
In this blog post, we discuss how AWS services were used to accelerate the development of a fast and sophisticated gene-editing prediction engine. We also describe our challenges and how AWS services helped us cost-effectively and efficiently navigate them, particularly with regard to handling large-scale datasets in a highly research-oriented scientific domain, while allowing us to retain flexibility.
About NRGene and CRISPR-IL
NRGene’s AI-based solutions are built around today’s next-generation sequencing technologies (NGS) and allow their analysis in an affordable, scalable, and precise manner. By applying our vast proprietary databases and algorithms, we provide leading international agriculture, seed development, and food companies with computational tools to optimize and accelerate their breeding programs, which significantly increase crop quality and save time and money.
At its core, the CRISPR-IL consortium is built around an iterative learning process, whereby researchers from multiple organizations design and carry-out CRISPR experiments on various organisms and cell types. GOGENOME measures the outcomes of these experiments and sends these results as feedback to a predictive engine that learns the most effective design for the next round of experiments. This predictive engine uses machine-learned models, using both publicly available data and proprietary data created inside the consortium. We train both feature-based and neural-network models and apply state of the art technologies, such as transfer-learning to reuse trained models across different organisms and cell-types.
CRISPR-IL’s infrastructure key challenges and initial requirements
We identified five key challenges in setting up the computational infrastructure needed for CRISPR-IL to succeed.
1. Organization challenges
The consortium comprises multiple members, across different research institutions and commercial companies, each having unique skills and interests. Facilitating collaboration and allowing people access to share information, data, models, services, and results in a secure yet hassle-free manner was a key consideration.
2. Risk mitigations challenges
The costs of undertaking CRISPR experiments are high, especially with the volume of clinical data the experiments generate. We realized a significant amount of infrastructure would need to be set up before any real experimental value could be evaluated for us to benefit from. Therefore, it was of the highest importance that a “minimum viable service” be set up as quickly as possible. This would allow us to produce experimental data and evaluate its utility and quality, before “burning” too much financial resources. Building the right environment quickly was a critical criterion to mitigate the risk of spending a lot of cash in futile and fruitless directions.
3. Chasing a “moving target”
CRISPR is an area of intense worldwide focus, and is rapidly evolving and growing. Scientifically, new enzymes are constantly discovered and new technologies are rapidly developed such as base-editing and prime-editing. Data-wise, more and more studies, which include high-throughput experiments are published and constantly submitted to public repositories. The CRISPR applications are now applied to an ever growing number of organisms and genomes, which should be stored, indexed and handled, with more and more data constantly submitted to public repositories while alternative approaches are constantly pursued and worked on. As a result, the architecture we would build – both for training our learning models and for serving predictions – would need to be flexible. Yet, it needed to be robust enough to incorporate these growing and changing datasets and approaches.
4. Navigating within the “fog of research”
Given the difficulty and complexity of the fundamental task at hand, designing and building a CRISPR gene-editing prediction engine, the computational environment and the service’s architecture had to be built to be amenable to small, incremental changes. It also had to allow the testing of research code “snippets” and ad-hoc computational modules, and to correctly incorporate different models into a single coherent system.
5. Challenges with scale
With CRISPR-IL being a research project trying to push the state-of-the-art, the level of uncertainty around the scale and exact computational needs of the consortium was high. We needed to build something that could easily scale up or down, and be able to change directions rather quickly to accommodate for the different needs that would arise once embarking on this project.
The system’s initial requirements
There were several factors to consider as we built the platform and made design decisions. We have fluctuations in resource requirements over time, with periodic peak demands that are orders of magnitude greater than the base-level day-to-day usage, but the needs can be hard to predict ahead of time. For example, we needed a platform that could automatically and quickly scale up and down to manage a variable volume of genomic sequencing data. In addition, the system needed to be flexible and reproducible for iterative scientific experiments, so we could quickly run the same experiment without manual overhead. We also wanted a machine learning framework that researchers would feel comfortable with, preferably one they are already somewhat familiar with.
We needed a pipeline that included having specific bioinformatic workflows process the results of experiments in preparation for machine learning. It had to be consistent with internal REST API servers to allow easy access to the data internally, for researchers to easily use in their day-to-day activities. We also needed an outward facing web application, to allow access to the final prediction engine externally, over the internet. In terms of data access, our architecture needed to support multiple organizations that collaborate around a single computational and data-storage platform.
Another aspect of consideration was our diverse set resource needs, including:
- Standalone compute environments for prototyping and research
- Elastic file storage
- Metadata data stores
- Inward facing and outward facing web servers
- Bioinformatic data pipelines
- ML training and serving
Each organization would handle one or more aspects of the overall process: some would do the actual experiments, others analyze the data and make it more understandable, and still others work on developing the ML models used for subsequent predictions. We also had to consider competing interests among participants, meaning each one should only have access to the specific and partial views he or she is authorized to see. Security and authorization mechanisms had to be used to verify that the different datasets or services are only available to the correct users. Finally, we needed to do all of this with a fast ramp-up with the initial infrastructure in place to provide ample time for the actual iterative research.
Key CRISPR-IL services built on top of AWS
To help us tackle all of our challenges and requirements, it seemed clear to us that AWS would be a great fit. Collaboration could be done across different organizations, with no overly complex networking for us to set up. We could just use the robust security building-blocks offered by AWS its extensive list of services in order to ramp up quickly. The elastic nature of AWS resources – storage and compute in particular – could help us mitigate risks around cost and scale. We could create different research environments for different people, catering to their idiosyncratic needs. Finally, we could build a data lake with a loosely coupled, distributed, microservice-oriented service that allowed for iterative changes coming from different research groups, synthesizing these contributions into a single holistic environment.
At the heart of the CRISPR-IL’s computational framework lies a web-server that performs the actual predictions and serves their results to users. We’ve named it: GoGenome. We built it using a wide range of AWS services, including Amazon S3 as the foundation of the data lake, Amazon EFS for shared file storage, Amazon RDS as the database engine, AWS Batch for data processing, and Amazon SageMaker for modeling and learning. In addition, GoGenome uses Amazon EKS to orchestrate the different microservices and models.
Our storage architecture can be summarized as such:
- Large amounts of raw sequencing data (>20 TB) are stored in Amazon S3.
- Amazon S3 also stores hundreds of millions of feature vectors, spanning several hundred gigabytes of tabular data.
- We use S3 Intelligent-Tiering where applicable, in addition to versioning to guarantee data is not written over or deleted accidentally.
- Using AWS Lambda events, we’ve also created an automated and secure uploading mechanism for our external sequencing centers. It includes authentication, object-tagging, and metadata entry insertions into an Amazon RDS database.
- Alongside Amazon S3, Amazon EFS is used to speed up access to genomic fasta files and their indexes as created by samtools, blast, tabix, minimap, and BWA.
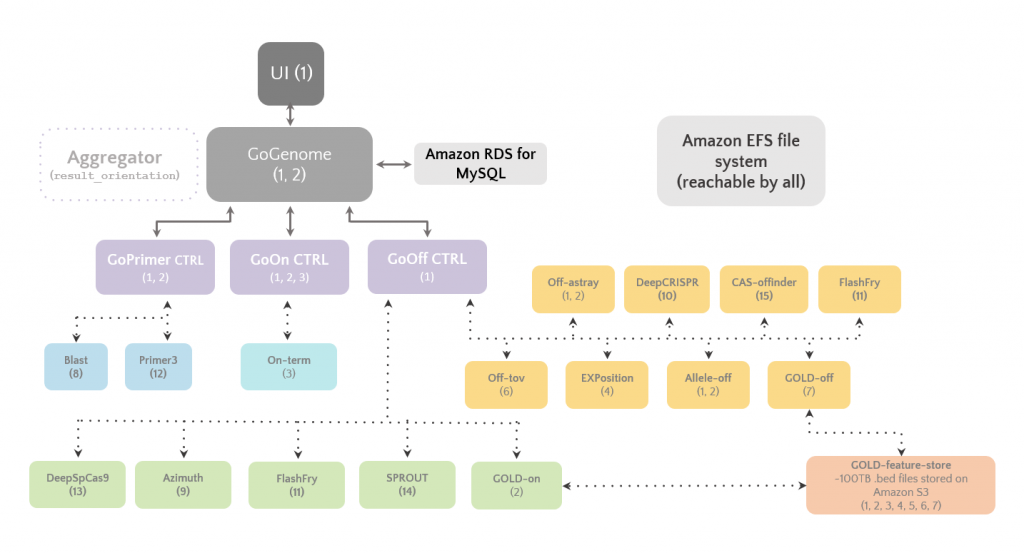
A schematic overview of GoGenome and the microservices powering it
1. NRGene (link)
2. Evogene (link)
3. Prof. Itay Mayrose, Tel-Aviv University (link)
4. Prof. Tamir Tuller, Tel-Aviv University (link)
5. Dr. David (Dudu) Burstein, Tel-Aviv University (link)
6. Dr. Isana Veksler, Ben-Gurion University (link)
7. Dr. Yaron Orenstein, Ben-Gurion University (link)
8. Blast (link) (source)
9. Azimuth (link) (source)
10. DeepCrispr (link) (source)
11. FlashFry (link) (source)
12. Primer3 (link) (source)
13. DeepSPCas9 (link) (source)
14. SPROUT (link) (source)
15. CAS-OFFinder (link) (source)
A second key service we’ve built on top of AWS is a high-throughput feature-store. It uses Amazon S3 and streamlines the data querying needed for learning and prediction, using an indexing mechanism spanning genomic regions of interests (ROIs). We’ve named the service: GOLD feature-store (GOLD is a reversed acronym of Deep Learning of GoGenome). GOLD-feature-store is both inexpensive to operate and sufficiently fast, making use of pre-computed and pre-indexed tables spanning billions of rows. It achieves sub-second latency that is adequate for our current needs, providing developers and researchers with quick access to feature-sets they require to train feature-based predictive models.

A schematic overview of the genomic GOLD-feature-store
CRISPR-IL’s full AWS tech stack
- Amazon S3 to store the large amounts of data needed for the prediction models.
- Amazon SageMaker for machine learning and serving of models.
- Amazon EKS to manage the various web services.
- Amazon EFS for fast read-access to large reference files across multiple microservices.
- AWS Batch for bioinformatic pipelines (based on Nextflow).
- Amazon RDS for MySQL for metadata storage.
- Amazon ECR for the Docker images we create, seamlessly integrated with GitHub actions for CI/CD.
- API Gateway for authentication and authorization.
- Workspaces for researchers to access the VPC without requiring VPNs or large data transfers.
- Infrastructure as code (using Terraform) to help manage an agile lifecycle of infrastructure, fast migration, and cloning to new AWS environments.
Summary of benefits
AWS enabled us to quickly build a robust and scalable computational framework that caters to the varying needs of multiple stakeholders: ML researchers, clinical researchers, computational biologists, and plant breeders, across multiple organizations and institutes. AWS helped us to scale seamlessly and separate development from production environments, allowing us to further develop the system while not disrupting the work of experiments, continuously deploy services from development to production, monitor the activity of services and users, control the running-costs of the different components, and do all these things securely and efficiently.
Thanks for taking some time to learn about NRGene, CRISPR-IL, and our journey with AWS. If you have any comments or questions about any of the AWS services uses, feel free to leave a comment in the comments section.
CRISPR-IL’s computational infrastructure is the product of the hard work done by many people, across multiple organizations:
- Dr. Eyal and the team at Evogene
- Dr. Mayrose’s, Dr. Burstein’s and Prof. Tuller’s labs at Tel-Aviv University
- Dr. Veksler-Lublinsky’s and Dr. Orenstein’s labs at Ben-Gurion University
- The R&D team at NRGene
The content and opinions in this post are those of the third-party author and AWS is not responsible for the content or accuracy of this post.