AWS Partner Network (APN) Blog
How Tamr Optimized Amazon EMR Workloads to Unify 200 Billion Records 5x Faster than On-Premises
By Clint Richardson, Technical Lead, Professional Services – Tamr
By Kalen Zhang, Partner Solutions Architect, Data & Analytics – AWS
By Marina Novikova, Startups Partner Solutions Architect – AWS
 |
| Tamr |
 |
Global business leaders recognize the value of advanced and augmented big data analytics over various internal and external data sources. However, technical leaders also face challenges capturing insights from data silos without unified master data.
Building up an elastic and scalable data unification solution and managing large-scale production environments across clusters brings extra engineering and operational challenges to organizations.
Tamr is an AWS ISV Partner with the Competencies in Machine Learning and Data and Analytics that helps enterprise customers migrate on-premises data unification workloads to Amazon Web Services (AWS) to take advantage of scalable and cost-effective managed services.
With Tamr, the artificial intelligence (AI)-powered data mastering solution matches and deduplicates the data to provide a new curated view of the underlying logical entities. These cleansed and mastered datasets are eventually fed into downstream reporting and analytics applications.
In this post, we’ll describe how migrating Tamr’s data mastering solutions from on-premises to AWS allowed a customer to process billions of records five times faster with fully managed Amazon EMR clusters.
In this example, migrating the customer to EMR was simple. Unfortunately, our first unoptimized run resulted in a processing time similar to on-premises (three weeks vs. four weeks).
After optimization of Amazon EMR, Apache HBase, and Apache Spark, however, Tamr was able to unify 200 billion records more than five times faster than performing the same tasks on-premises, thereby enabling the customer to achieve analytics at scale in AWS.
Challenges
A life sciences customer had challenges unifying its real-world healthcare data using on-premises Hadoop clusters in a timeline under several weeks. This resulted in minimized data consumption from analysts and prevented cross-source insights into how drugs and medical products perform in the real world.
The business and technical challenges for unifying 200 billion records of real-world data are inextricably connected and boil down to one core metric: the speed of processing.
On the business side, it’s hard to build buy-in for a new data initiative if consumers have to spend weeks waiting for the data to arrive in a usable format after every delivery.
On the technical side, doing O(100) non-linear operations on the scale of hundreds of billions of records requires a highly optimized architecture and execution engine to deliver the data at all, let alone quickly.
Implementation
Using Tamr’s AWS footprint, we were able to leverage multiple big data technologies to solve the technical challenges and in turn drive business adoption and empowerment.
Tamr migrated on-premises data unification workloads to Amazon EMR, tuned EMR clusters, and achieved more than a five times speedup in processing 200 billion records, providing answers in days not weeks.
Ease of Migration to Amazon EMR
Migrating our workflow to Amazon EMR services was a straightforward process. EMR makes available nearly all flavors and versions of Hadoop components needed, along with the assurances they’ll work together from a networking and configuration authentication point of view. With Tamr’s modular dependency architecture, migrating from the world of on-premises to AWS was as easy as plug-and-play.
Additionally, we had the benefit of being able to scale our AWS footprint as needed, taking advantage of managed Apache HBase and Apache Spark on Amazon EMR clusters to spin up massive amounts of compute when needed and turn them back off to save costs when no longer used.
This ability became critically important during the performance tuning phase, especially for allowing per-job tuning so every single job could be run in a performant way depending on its data access and computation patterns.
Tuning Amazon EMR
The initial attempt at migrating to AWS scaled not only the number of nodes in the EMR HBase cluster, but also the number of tables. The data would be represented as multiple—O(100)—datasets inside Tamr’s processing.
Our initial thought was to separate these out as different tables inside of HBase (that is, each dataset would get its own dedicated HBase table). That way, the tables themselves would be well distributed among the region servers (see Figure 1). Thus, when we ran against these tables with multiple, 1,000-virtual core Apache Spark on Amazon EMR clusters concurrently, no single region server would be too bottlenecked.
Unfortunately, this approach did not work, and we saw severely slow HBase read performance throughout the pipeline execution. This slowdown caused the full processing time to come in just under one month during our first run after moving to AWS.
The biggest driver in this was HBase cold-spotting, with some region servers taking inordinate amounts of time to pull the data from Amazon Simple Storage Service (Amazon S3) even though they did not show a large load.
Figure 1 – Tamr datasets distributed across HBase region servers as separate tables.
After Optimization
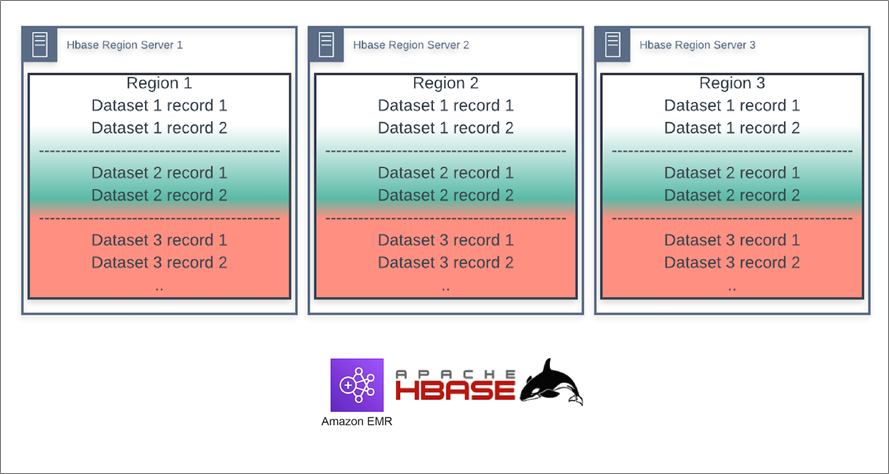
A more successful approach was to simply put all datasets (yes, all 200 billion records) into one HBase table. We salted the row-keys to ensure every dataset was uniformly distributed across the single table to avoid hot-spotting.
The key from a performance point of view is to make the number of salt values equal to the number of regions in your HBase table (or a multiple of it) so that your Spark tasks are pulling from different regions (see Figure 2). Using a random hash as the salt value ensures rows are distributed evenly across the different HBase regions for all datasets.
Figure 2 – Tamr datasets distributed across HBase region servers as one table.
Lessons Learned
The following four points are key to the success of the optimization described:
1. Improving Spark Performance with Amazon S3
When writing the data in Parquet out to Amazon S3, use EMRFS with its S3-optimized committer, rather than Spark’s s3a driver. This change easily cut more than a day off our processing time when summing over all our exports, effectively speeding up our second attempt by 30%.
The underlying cause stemmed from tension between how we want to process the data and how we want to store it.
For processing, we want thousands of partitions to take advantage of the large EMR clusters at our fingertips. However, for storage we don’t want thousands of files. Because Spark by default writes one or more temporary files per partition, which then need to be combined, using the default s3a committer resulted in extremely slow performance for the combine and cleanup stage (where many files need to be shuffled around S3).
Learn more about improving Spark performance with Amazon S3.
2. Strategizing HBase Table Design
Separating each dataset as its own table was the wrong way to scale. We had initially tried this approach because each dataset corresponded to different source records that would have different read and write patterns as we unified the data.
Consolidating each dataset into a single table with row-key salting was a better method. The following table shows the differences in how we stored datasets in HBase.
| Initial run | Optimized run | |
| Number of Tamr datasets | 80 | 80 |
| Number of HBase tables | 80 | 1 |
| Regions per table | 120 | 5,400 |
| Total HBase regions | 10,800 | 5,400 |
| Salt values per table | 1,000 | 5,400 |
| Number of HBase region servers | 180 | 180 |
| HBase region server machine type | r5.xlarge | r5.xlarge |
A key point of this optimization is that, in the initial run, we configured each Tamr dataset the same even though they had vastly different scales. For example, some hold hundreds of thousands of records, while some hold tens of billions. This meant the largest tables underwent multiple splits while the smallest had an inefficient distribution of regions and salt values.
We could have optimized each table individually, but simply putting them together with a suitable salting scheme allowed us to take advantage of the statistical distribution of records and achieve consistent and considerable performance.
3. Optimizing HBase Regions
We also hadn’t done a fine enough granularity optimization of regions and their sizes for the different tables. We initially set the region size to be 10 GB, but this again led to larger tables undergoing multiple splits and small tables being inefficiently stored.
After the initial run, we simply counted the amount of data stored during processing (~100 TB) and then divided this by the total number of regions to settle on 20GB per region (see the following table).
In truth, both of these numbers could have been changed to get us to a total storage of ~100 TB. We settled on this distribution because more regions would have meant more Spark tasks. And, given we were spinning up ~ 1,000 vCore Spark clusters, we didn’t want any single Spark stage to have more than five times more tasks than cores.
| Configuration parameter | Initial run | Optimized run |
| hbase.hregion.max.filesize | 10737418240 | 21474836480 |
4. Further Tuning Spark Clusters
One major advantage we had operating in AWS vs. on-premises is that we had the ability to spin up massive amounts of concurrent, ephemeral compute as needed during processing.
The overall processing graph greatly benefited from the use of independent EMR clusters. We also had the ability to use whatever machine types were needed for that job. We typically used clusters containing 40 r5.8xlarge nodes due to their higher Amazon Elastic Block Store (Amazon EBS) and networking speeds.
To take advantage of the changes we implemented in Apache HBase, our Spark settings need to be consistent with both points 2 and 3 above. There are effectively the same number of concurrently runnable Spark tasks as there are salt-key values. This means if your tables have 1,000 salt values, then effectively only 1,000 Spark tasks can run in parallel, regardless of how many cores you have.
This requirement only affects the reading phase, as later data manipulations will start with the number of output partitions from the previous stage. Generally, it’s simpler to align these around a reasonable value.
Because we were using 5,400 salt values in HBase, but were doing significant data manipulation that would benefit from smaller partitions during the processing stage, we settled on the Spark configuration parameters shown in the following table. Note that Tamr makes use of both the Spark DataFrame and lower-level RDD APIs, which is why we set both SQL partitions and shuffle parallelism.
| Configuration parameter | Initial run | Optimized run |
| spark.sql.shuffle.partitions | 2,200 | 10,000 |
| spark.default.parallelism | 1,800 | 9,000 |
| spark.checkpoint.compress | false | true |
| spark.io.compression.codec | n/a | snappy |
| spark.io.compression.snappy.blockSize | n/a | 128,000 |
| spark.rdd.compress | false | true |
| spark.shuffle.spill.compress | false | true |
| spark.memory.storageFraction | 0.5 | 0.0 |
Outcomes
The above performance tuning allowed us to run multiple concurrent EMR Spark clusters, all reading from the same EMR HBase cluster to unify hundreds of billions of records in 2.5 days, whereas on-premises processing took more than two weeks.
Conclusion
Tamr’s cloud-native data mastering solution enables enterprises to accelerate big data analytics using Amazon EMR clusters.
Running Tamr’s data mastering solution on AWS allows organizations to unify hundreds of billions of records in days by modernizing their approach to data management with fully managed Amazon EMR clusters.
As a result, organizations can easily and quickly break down data silos and capture insights to increase revenue, reduce risk, and improve customer service. Meet with Tamr to learn more about how Tamr helps enterprises connect data sources to drive business.
Tamr – AWS Partner Spotlight
Tamr is an AWS Competency Partner that helps enterprise customers migrate on-premises data unification workloads to AWS to take advantage of scalable and cost-effective managed services.
Contact Tamr | Partner Overview | AWS Marketplace
*Already worked with Tamr? Rate the Partner
*To review an AWS Partner, you must be a customer that has worked with them directly on a project.